-
 Specific Antimicrobial Activities Revealed by Comparative Evaluation of Selected Gemmotherapy Extracts
Specific Antimicrobial Activities Revealed by Comparative Evaluation of Selected Gemmotherapy Extracts -
 Comparison of Antibiotic Susceptibility Testing Results for Gram-Positive Mastitis Pathogens from Dairy Cows
Comparison of Antibiotic Susceptibility Testing Results for Gram-Positive Mastitis Pathogens from Dairy Cows -
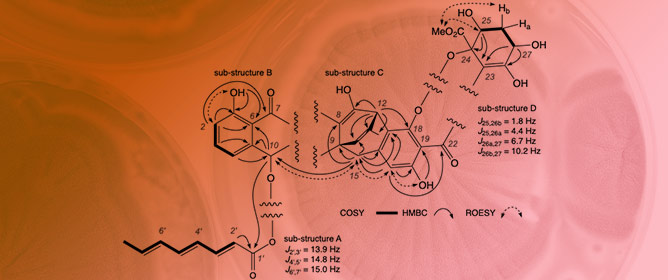 Jugiones A–D: Antibacterial Xanthone–Anthraquinone Heterodimers from Australian Soil-Derived Penicillium shearii CMB-STF067
Jugiones A–D: Antibacterial Xanthone–Anthraquinone Heterodimers from Australian Soil-Derived Penicillium shearii CMB-STF067
Journal Description
Antibiotics
Antibiotics
is an international, peer-reviewed, open access journal on all aspects of antibiotics, published monthly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, Embase, CAPlus / SciFinder, and other databases.
- Journal Rank: JCR - Q1 (Pharmacology & Pharmacy) / CiteScore - Q1 (General Pharmacology, Toxicology and Pharmaceutics)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 13.7 days after submission; acceptance to publication is undertaken in 2.5 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
4.8 (2022);
5-Year Impact Factor:
4.9 (2022)
Latest Articles
The Genetic Landscape of Antimicrobial Resistance Genes in Enterococcus cecorum Broiler Isolates
Antibiotics 2024, 13(5), 409; https://doi.org/10.3390/antibiotics13050409 (registering DOI) - 29 Apr 2024
Abstract
Enterococcus cecorum is associated with bacterial chondronecrosis with osteomyelitis (BCO) in broilers. Prophylactic treatment with antimicrobials is common in the poultry industry, and, in the case of outbreaks, antimicrobial treatment is needed. In this study, the minimum inhibitory concentrations (MICs) and epidemiological cutoff
[...] Read more.
Enterococcus cecorum is associated with bacterial chondronecrosis with osteomyelitis (BCO) in broilers. Prophylactic treatment with antimicrobials is common in the poultry industry, and, in the case of outbreaks, antimicrobial treatment is needed. In this study, the minimum inhibitory concentrations (MICs) and epidemiological cutoff (ECOFF) values (COWT) for ten antimicrobials were determined in a collection of E. cecorum strains. Whole-genome sequencing data were analyzed for a selection of these E. cecorum strains to identify resistance determinants involved in the observed phenotypes. Wild-type and non-wild-type isolates were observed for the investigated antimicrobial agents. Several antimicrobial resistance genes (ARGs) were detected in the isolates, linking phenotypes with genotypes for the resistance to vancomycin, tetracycline, lincomycin, spectinomycin, and tylosin. These detected resistance genes were located on mobile genetic elements (MGEs). Point mutations were found in isolates with a non-wild-type phenotype for enrofloxacin and ampicillin/ceftiofur. Isolates showing non-wild-type phenotypes for enrofloxacin had point mutations within the GyrA, GyrB, and ParC proteins, while five amino acid changes in penicillin-binding proteins (PBP2x superfamily) were observed in non-wild-type phenotypes for the tested β-lactam antimicrobials. This study is one of the first that describes the genetic landscape of ARGs within MGEs in E. cecorum, in association with phenotypical resistance determination.
Full article
(This article belongs to the Special Issue Antibiotics Resistance in Animals and the Environment)
Open AccessArticle
Adsorption of Macrolide Antibiotics and a Metabolite onto Polyethylene Terephthalate and Polyethylene Microplastics in Aquatic Environments
by
Carmen Mejías, Julia Martín, Laura Martín-Pozo, Juan Luis Santos, Irene Aparicio and Esteban Alonso
Antibiotics 2024, 13(5), 408; https://doi.org/10.3390/antibiotics13050408 (registering DOI) - 29 Apr 2024
Abstract
Microplastics (MPs) and antibiotics are emerging pollutants widely found in aquatic environments, potentially causing environmental harm. MPs may act as carriers for antibiotics, affecting their environmental distribution. This study investigates the adsorption of four macrolide antibiotics and a metabolite onto two types of
[...] Read more.
Microplastics (MPs) and antibiotics are emerging pollutants widely found in aquatic environments, potentially causing environmental harm. MPs may act as carriers for antibiotics, affecting their environmental distribution. This study investigates the adsorption of four macrolide antibiotics and a metabolite onto two types of MPs: polyethylene terephthalate (PET) and polyethylene (PE). Results revealed a linear isotherm adsorption model, with higher adsorption to PET than to PE (R2 > 0.936 for PE and R2 > 0.910 for PET). Hydrophobic interactions and hydrogen bonding could be the main adsorption mechanisms, with pore filling potentially involved. Reduced particle size enhances adsorption due to the increase of active adsorption sites. This increasement is more pronounced in PE than in PET, leading to an 11.6% increase in the average adsorption of all macrolides to PE, compared to only 5.1% to PET. Dissolved organic matter inhibits adsorption (azithromycin adsorption to PE was reduced from 12% to 5.1%), while salinity enhances it just until 1% salinity. pH slightly influences adsorption, with maximal adsorption at neutral pH. Results in real samples showed that complexity of the matrix decreased adsorption. Overall, these findings indicate that PE and PET MPs can be a vector of macrolides in aquatic environments.
Full article
(This article belongs to the Special Issue Environmental Fate and Effects of Antibiotics and Antibiotic Resistance Genes)
Open AccessArticle
Fungal Bioremediation of the β-Lactam Antibiotic Ampicillin under Laccase-Induced Conditions
by
Bouthaina Ghariani, Abdulrahman H. Alessa, Imen Ben Atitallah, Ibtihel Louati, Ahmad A. Alsaigh, Tahar Mechichi and Héla Zouari-Mechichi
Antibiotics 2024, 13(5), 407; https://doi.org/10.3390/antibiotics13050407 (registering DOI) - 29 Apr 2024
Abstract
Due to widespread overuse, pharmaceutical compounds, such as antibiotics, are becoming increasingly prevalent in greater concentrations in aquatic ecosystems. In this study, we investigated the capacity of the white-rot fungus, Coriolopsis gallica (a high-laccase-producing fungus), to biodegrade ampicillin under different cultivation conditions. The
[...] Read more.
Due to widespread overuse, pharmaceutical compounds, such as antibiotics, are becoming increasingly prevalent in greater concentrations in aquatic ecosystems. In this study, we investigated the capacity of the white-rot fungus, Coriolopsis gallica (a high-laccase-producing fungus), to biodegrade ampicillin under different cultivation conditions. The biodegradation of the antibiotic was confirmed using high-performance liquid chromatography, and its antibacterial activity was evaluated using the bacterial growth inhibition agar well diffusion method, with Escherichia coli as an ampicillin-sensitive test strain. C. gallica successfully eliminated ampicillin (50 mg L−1) after 6 days of incubation in a liquid medium. The best results were achieved with a 9-day-old fungal culture, which treated a high concentration (500 mg L−1) of ampicillin within 3 days. This higher antibiotic removal rate was concomitant with the maximum laccase production in the culture supernatant. Meanwhile, four consecutive doses of 500 mg L−1 of ampicillin were removed by the same fungal culture within 24 days. After that, the fungus failed to remove the antibiotic. The measurement of the ligninolytic enzyme activity showed that C. gallica laccase might participate in the bioremediation of ampicillin.
Full article
(This article belongs to the Special Issue Environmental Fate of Antibiotics: Monitoring, Toxicity, Resistance, and Removal Methods)
►▼
Show Figures
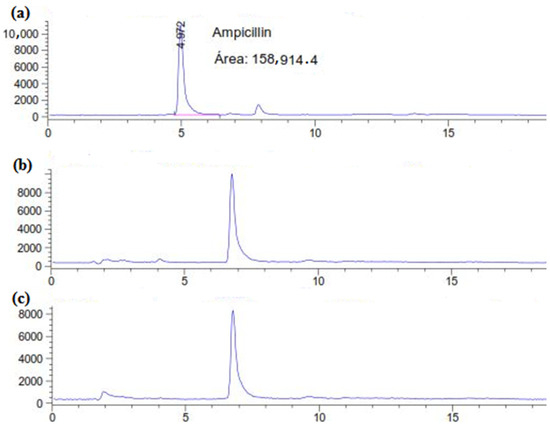
Figure 1
Open AccessReview
The Interplay between Antibiotics and the Host Immune Response in Sepsis: From Basic Mechanisms to Clinical Considerations: A Comprehensive Narrative Review
by
Martina Tosi, Irene Coloretti, Marianna Meschiari, Sara De Biasi, Massimo Girardis and Stefano Busani
Antibiotics 2024, 13(5), 406; https://doi.org/10.3390/antibiotics13050406 (registering DOI) - 28 Apr 2024
Abstract
Sepsis poses a significant global health challenge due to immune system dysregulation. This narrative review explores the complex relationship between antibiotics and the immune system, aiming to clarify the involved mechanisms and their clinical impacts. From pre-clinical studies, antibiotics exhibit various immunomodulatory effects,
[...] Read more.
Sepsis poses a significant global health challenge due to immune system dysregulation. This narrative review explores the complex relationship between antibiotics and the immune system, aiming to clarify the involved mechanisms and their clinical impacts. From pre-clinical studies, antibiotics exhibit various immunomodulatory effects, including the regulation of pro-inflammatory cytokine production, interaction with Toll-Like Receptors, modulation of the P38/Pmk-1 Pathway, inhibition of Matrix Metalloproteinases, blockade of nitric oxide synthase, and regulation of caspase-induced apoptosis. Additionally, antibiotic-induced alterations to the microbiome are associated with changes in systemic immunity, affecting cellular and humoral responses. The adjunctive use of antibiotics in sepsis patients, particularly macrolides, has attracted attention due to their immune-regulatory effects. However, there are limited data comparing different types of macrolides. More robust evidence comes from studies on community-acquired pneumonia, especially in severe cases with a hyper-inflammatory response. While studies on septic shock have shown mixed results regarding mortality rates and immune response modulation, conflicting findings are also observed with macrolides in acute respiratory distress syndrome. In conclusion, there is a pressing need to tailor antibiotic therapy based on the patient’s immune profile to optimize outcomes in sepsis management.
Full article
(This article belongs to the Special Issue Antibacterial Resistance and Infection Control in ICU)
►▼
Show Figures
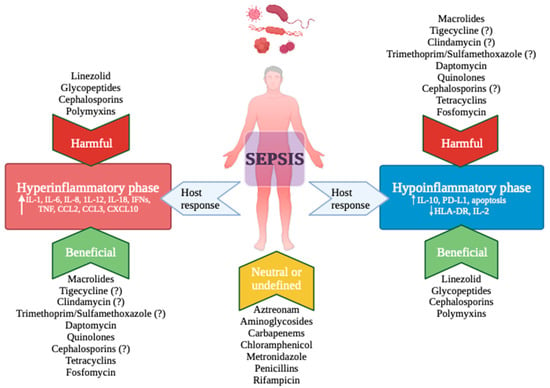
Figure 1
Open AccessArticle
Antifungal Activity of Brilacidin, a Nonpeptide Host Defense Molecule
by
David J. Larwood and David A. Stevens
Antibiotics 2024, 13(5), 405; https://doi.org/10.3390/antibiotics13050405 (registering DOI) - 28 Apr 2024
Abstract
Natural host defensins, also sometimes termed antimicrobial peptides, are evolutionarily conserved. They have been studied as antimicrobials, but some pharmaceutical properties, undesirable for clinical use, have led to the development of synthetic molecules with constructed peptide arrangements and/or peptides not found in nature.
[...] Read more.
Natural host defensins, also sometimes termed antimicrobial peptides, are evolutionarily conserved. They have been studied as antimicrobials, but some pharmaceutical properties, undesirable for clinical use, have led to the development of synthetic molecules with constructed peptide arrangements and/or peptides not found in nature. The leading development currently is synthetic small-molecule nonpeptide mimetics, whose physical properties capture the characteristics of the natural molecules and share their biological attributes. We studied brilacidin, an arylamide of this type, for its activity in vitro against fungi (40 clinical isolates, 20 species) that the World Health Organization has highlighted as problem human pathogens. We found antifungal activity at low concentrations for many pathogens, which indicates that further screening for activity, particularly in vivo, is justified to evaluate this compound, and other mimetics, as attractive leads for the development of effective antifungal agents.
Full article
Open AccessArticle
Broad-Spectrum In Vitro Activity of Nα-aroyl-N-aryl-Phenylalanine Amides against Non-Tuberculous Mycobacteria and Comparative Analysis of RNA Polymerases
by
Markus Lang, Uday S. Ganapathy, Rana Abdelaziz, Thomas Dick and Adrian Richter
Antibiotics 2024, 13(5), 404; https://doi.org/10.3390/antibiotics13050404 (registering DOI) - 28 Apr 2024
Abstract
This study investigates the in vitro activity of Nα-aroyl-N-aryl-phenylalanine amides (AAPs), previously identified as antimycobacterial RNA polymerase (RNAP) inhibitors, against a panel of 25 non-tuberculous mycobacteria (NTM). The compounds, including the hit compound MMV688845, were selected based on their structural
[...] Read more.
This study investigates the in vitro activity of Nα-aroyl-N-aryl-phenylalanine amides (AAPs), previously identified as antimycobacterial RNA polymerase (RNAP) inhibitors, against a panel of 25 non-tuberculous mycobacteria (NTM). The compounds, including the hit compound MMV688845, were selected based on their structural diversity and previously described activity against mycobacteria. Bacterial strains, including the M. abscessus complex, M. avium complex, and other clinically relevant NTM, were cultured and subjected to growth inhibition assays. The results demonstrate significant activity against the most common NTM pathogens from the M. abscessus and M. avium complexes. Variations in activity were observed against other NTM species, with for instance M. ulcerans displaying high susceptibility and M. xenopi and M. simiae resistance to AAPs. Comparative analysis of RNAP β and β′ subunits across mycobacterial species revealed strain-specific polymorphisms, providing insights into differential compound susceptibility. While conservation of target structures was observed, differences in compound activity suggested influences beyond drug–target interactions. This study highlights the potential of AAPs as effective antimycobacterial agents and emphasizes the complex interplay between compound structure, bacterial genetics, and in vitro activity.
Full article
(This article belongs to the Special Issue Therapeutic and Microbiological Approaches for Combating Non-tuberculous Mycobacterial Infections)
Open AccessArticle
Antimicrobial Use in Pig Farms in the Midwestern Region of Minas Gerais, Brazil
by
Bruno César de Oliveira, Idael Christiano de Almeida Santa Rosa, Maurício Cabral Dutra, Felipe Norberto Alves Ferreira, Andrea Micke Moreno, Luisa Zanolli Moreno, Júlia da Mata Góes Silva, Simone Koprowski Garcia and Dalton de Oliveira Fontes
Antibiotics 2024, 13(5), 403; https://doi.org/10.3390/antibiotics13050403 (registering DOI) - 28 Apr 2024
Abstract
The use of antimicrobials in swine production is an issue that concerns the whole world due to their impact on animal and public health. This study aimed to verify the antimicrobial use in 29 commercial full-cycle farms in the midwestern region of the
[...] Read more.
The use of antimicrobials in swine production is an issue that concerns the whole world due to their impact on animal and public health. This study aimed to verify the antimicrobial use in 29 commercial full-cycle farms in the midwestern region of the state of Minas Gerais, since this region is a hub of intensive pig farming in Brazil, as well as the possible correlations between the use of antimicrobials, biosecurity, and productivity. A total of 28 different drugs used for preventive purposes were described. On average, the herds used seven drugs, exposing the piglets for 116 days and totaling 434.17 mg of antimicrobials per kilogram of pig produced. Just eight active ingredients made up 77.5% of the total number of drugs used on the studied herds. Significant differences were found between the variables, biosecurity score and number of sows, antimicrobial amount and number of drugs, number of drugs and number of sows, and between productivity and biosecurity scores. The use of antimicrobials was considered excessive in the swine farms in the state of Minas Gerais compared to what was reported in Brazil and in other countries. Educational measures and better control should be proposed to reduce the preventive use of antimicrobials.
Full article
(This article belongs to the Special Issue One Health and Antibiotic Use in Veterinary Medicine)
►▼
Show Figures
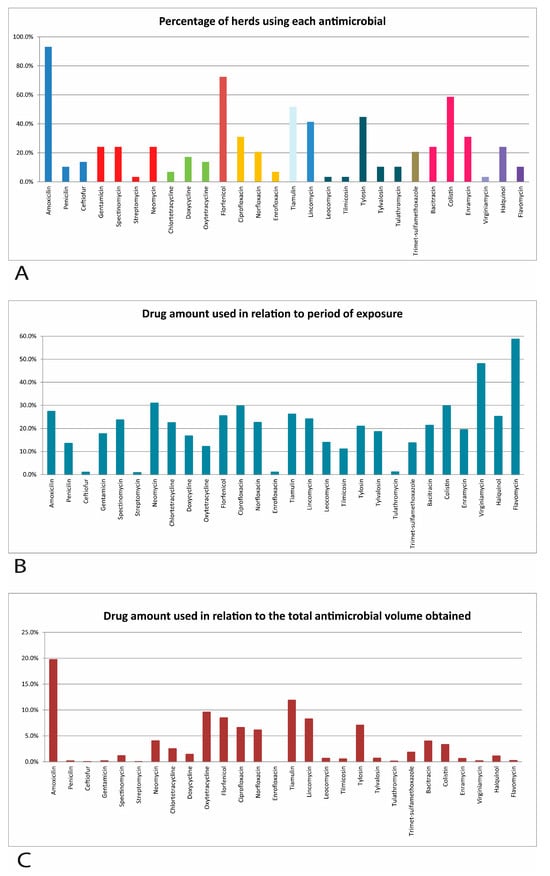
Figure 1
Open AccessEditorial
Molecular Characterization of Gram-Negative Bacteria: Antimicrobial Resistance, Virulence and Epidemiology
by
Theodoros Karampatakis
Antibiotics 2024, 13(5), 402; https://doi.org/10.3390/antibiotics13050402 (registering DOI) - 28 Apr 2024
Abstract
Multidrug-resistant (MDR), extensively drug-resistant (XDR) and pan-drug-resistant (PDR) Gram-negative bacteria constitute a huge public health problem [...]
Full article
(This article belongs to the Special Issue Molecular Characterization of Gram-Negative Bacteria: Antimicrobial Resistance, Virulence and Epidemiology)
Open AccessArticle
Characterisation of PVL-Positive Staphylococcus argenteus from the United Arab Emirates
by
Stefan Monecke, Sindy Burgold-Voigt, Sascha D. Braun, Celia Diezel, Elisabeth M. Liebler-Tenorio, Elke Müller, Rania Nassar, Martin Reinicke, Annett Reissig, Abiola Senok and Ralf Ehricht
Antibiotics 2024, 13(5), 401; https://doi.org/10.3390/antibiotics13050401 (registering DOI) - 27 Apr 2024
Abstract
Staphylococcus argenteus is a recently described staphylococcal species that is related to Staphylococcus aureus but lacks the staphyloxanthin operon. It is able to acquire both resistance markers such as the SCCmec elements and mobile genetic elements carrying virulence-associated genes from S. aureus
[...] Read more.
Staphylococcus argenteus is a recently described staphylococcal species that is related to Staphylococcus aureus but lacks the staphyloxanthin operon. It is able to acquire both resistance markers such as the SCCmec elements and mobile genetic elements carrying virulence-associated genes from S. aureus. This includes those encoding the Panton–Valentine leukocidin (PVL), which is associated mainly with severe and/or recurrent staphylococcal skin and soft tissue infections. Here, we describe the genome sequences of two PVL-positive, mecA-negative S. argenteus sequence type (ST) 2250 isolates from the United Arab Emirates in detail. The isolates were found in a dental clinic in the United Arab Emirates (UAE). Both were sequenced using Oxford Nanopore Technology (ONT). This demonstrated the presence of temperate bacteriophages in the staphylococcal genomes, including a PVL prophage. It was essentially identical to the published sequence of phiSa2wa_st78 (GenBank NC_055048), a PVL phage from an Australian S. aureus clonal complex (CC) 88 isolate. Besides the PVL prophage, one isolate carried another prophage and the second isolate carried two additional prophages, whereby the region between these two prophages was inverted. This “flipped” region comprised about 1,083,000 bp, or more than a third of the strain’s genome, and it included the PVL prophage. Prophages were induced by Mitomycin C treatment and subjected to transmission electron microscopy (TEM). This yielded, in accordance to the sequencing results, one or, respectively, two distinct populations of icosahedral phages. It also showed prolate phages which presumptively might be identified as the PVL phage. This observation highlights the significance bacteriophages have as agents of horizontal gene transfer as well as the need for monitoring emerging staphylococcal strains, especially in cosmopolitan settings such as the UAE.
Full article
(This article belongs to the Special Issue Staphylococcus— Molecular Pathogenesis, Virulence Regulation and Antibiotics Resistance)
►▼
Show Figures
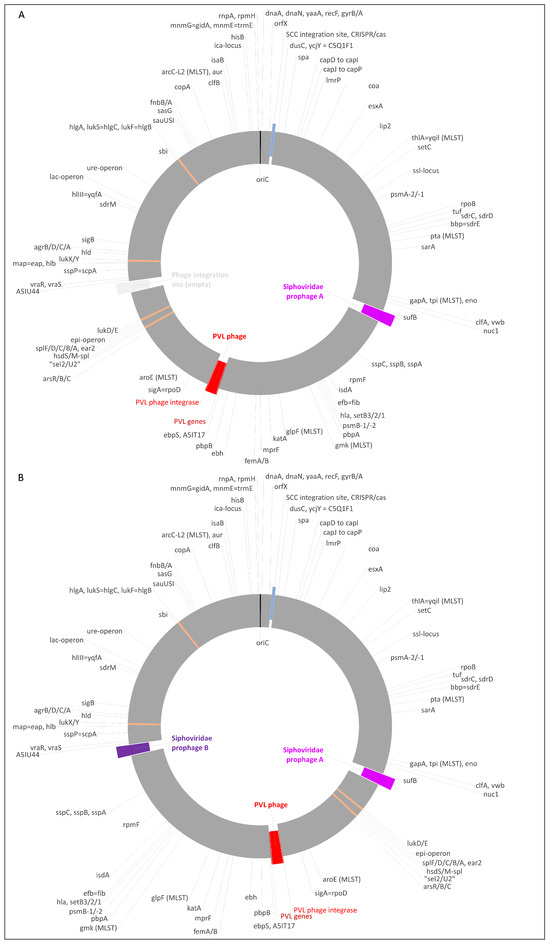
Figure 1
Open AccessArticle
Prevalence of Infections and Antimicrobial Resistance of ESKAPE Group Bacteria Isolated from Patients Admitted to the Intensive Care Unit of a County Emergency Hospital in Romania
by
Alina-Simona Bereanu, Rareș Bereanu, Cosmin Mohor, Bogdan Ioan Vintilă, Ioana Roxana Codru, Ciprian Olteanu and Mihai Sava
Antibiotics 2024, 13(5), 400; https://doi.org/10.3390/antibiotics13050400 (registering DOI) - 27 Apr 2024
Abstract
The ESKAPE group (Enterococcus faecium, Staphylococcus aureus, Klebsiella Pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, Enterobacter spp.) is a group of bacteria very difficult to treat due to their high ability to acquire resistance to antibiotics and are the
[...] Read more.
The ESKAPE group (Enterococcus faecium, Staphylococcus aureus, Klebsiella Pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, Enterobacter spp.) is a group of bacteria very difficult to treat due to their high ability to acquire resistance to antibiotics and are the main cause of nosocomial infections worldwide, posing a threat to global public health. Nosocomial infections with MDR bacteria are found mainly in Intensive Care Units, due to the multitude of maneuvers and invasive medical devices used, the prolonged antibiotic treatments, the serious general condition of these critical patients, and the prolonged duration of hospitalization. Materials and Methods: During a period of one year, from January 2023 to December 2023, this cross-sectional study was conducted on patients diagnosed with sepsis admitted to the Intensive Care Unit of the Sibiu County Emergency Clinical Hospital. Samples taken were tracheal aspirate, catheter tip, pharyngeal exudate, wound secretion, urine culture, blood culture, and peritoneal fluid. Results: The most common bacteria isolated from patients admitted to our Intensive Care Unit was Klebsiella pneumoniae, followed by Acinetobacter baumanii and Pseudomonas aeruginosa. Gram-positive cocci (Enterococcus faecium and Staphilococcus aureus) were rarely isolated. Most of the bacteria isolated were MDR bacteria. Conclusions: The rise of antibiotic and antimicrobial resistance among strains in the nosocomial environment and especially in Intensive Care Units raises serious concerns about limited treatment options.
Full article
(This article belongs to the Special Issue Antimicrobial Resistance and Epidemiological Study of Clinically Relevant Pathogens)
►▼
Show Figures
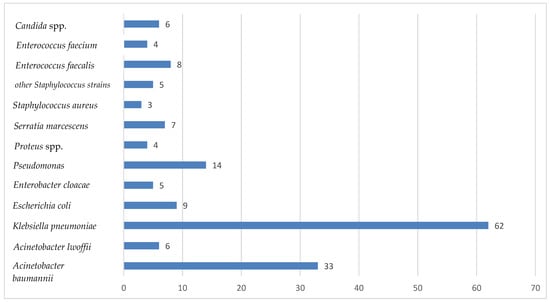
Figure 1
Open AccessArticle
Microbiology of Prosthetic Joint Infections: A Retrospective Study of an Italian Orthopaedic Hospital
by
Virginia Suardi, Daniele Baroni, Abdelrahman Hosni Abdelhamid Shahein, Valentina Morena, Nicola Logoluso, Laura Mangiavini and Antonio Virgilio Pellegrini
Antibiotics 2024, 13(5), 399; https://doi.org/10.3390/antibiotics13050399 - 26 Apr 2024
Abstract
The most frequent cause of periprosthetic infections (PJIs) is intraoperative contamination; hence, antibiotic prophylaxis plays a crucial role in prevention. Modifications to standard prophylaxis can be considered if there is a high incidence of microorganisms resistant to current protocols. To date, very few
[...] Read more.
The most frequent cause of periprosthetic infections (PJIs) is intraoperative contamination; hence, antibiotic prophylaxis plays a crucial role in prevention. Modifications to standard prophylaxis can be considered if there is a high incidence of microorganisms resistant to current protocols. To date, very few studies regarding microbial etiology have been published in Italy. In this single-center, retrospective study conducted at IRCCS Ospedale Galeazzi-Sant’Ambrogio in Milan, we analyzed hip, knee, and shoulder PJIs in patients undergoing first implantation between 1 January 17 and 31 December 2021. The primary aim was to derive a local microbiological case history. The secondary aim was to evaluate the adequacy of preoperative antibiotic prophylaxis in relation to the identified bacteria. A total of 57 PJIs and 65 pathogens were identified: 16 S. aureus, 15 S. epidermidis, and 10 other coagulase-negative staphylococci (CoNS), which accounted for 63% of the isolations. A total of 86.7% of S. epidermidis were methicillin-resistant (MRSE). In line with other case reports, we found a predominance of staphylococcal infections, with a lower percentage of MRSA than the Italian average, while we found a high percentage of MRSE. We estimated that 44.6% of the bacteria isolated were resistant to cefazolin, our standard prophylaxis. These PJIs could be prevented by using glycopeptide alone or in combination with cefazolin, but the literature reports conflicting results regarding the adequacy of such prophylaxis. In conclusion, our study showed that in our local hospital, our standard antibiotic prophylaxis is ineffective for almost half of the cases, highlighting the importance of defining specific antibiotic guidelines based on the local bacterial prevalence of each institution.
Full article
(This article belongs to the Special Issue Antimicrobial Therapeutics for Bone and Periprosthetic Joint Infection)
Open AccessArticle
Discovery of Antimicrobial Agents Based on Structural and Functional Study of the Klebsiella pneumoniae MazEF Toxin–Antitoxin System
by
Chenglong Jin, Sung-Min Kang, Do-Hee Kim, Yuno Lee and Bong-Jin Lee
Antibiotics 2024, 13(5), 398; https://doi.org/10.3390/antibiotics13050398 - 26 Apr 2024
Abstract
►▼
Show Figures
Klebsiella pneumoniae causes severe human diseases, but its resistance to current antibiotics is increasing. Therefore, new antibiotics to eradicate K. pneumoniae are urgently needed. Bacterial toxin–antitoxin (TA) systems are strongly correlated with physiological processes in pathogenic bacteria, such as growth arrest, survival, and
[...] Read more.
Klebsiella pneumoniae causes severe human diseases, but its resistance to current antibiotics is increasing. Therefore, new antibiotics to eradicate K. pneumoniae are urgently needed. Bacterial toxin–antitoxin (TA) systems are strongly correlated with physiological processes in pathogenic bacteria, such as growth arrest, survival, and apoptosis. By using structural information, we could design the peptides and small-molecule compounds that can disrupt the binding between K. pneumoniae MazE and MazF, which release free MazF toxin. Because the MazEF system is closely implicated in programmed cell death, artificial activation of MazF can promote cell death of K. pneumoniae. The effectiveness of a discovered small-molecule compound in bacterial cell killing was confirmed through flow cytometry analysis. Our findings can contribute to understanding the bacterial MazEF TA system and developing antimicrobial agents for treating drug-resistant K. pneumoniae.
Full article
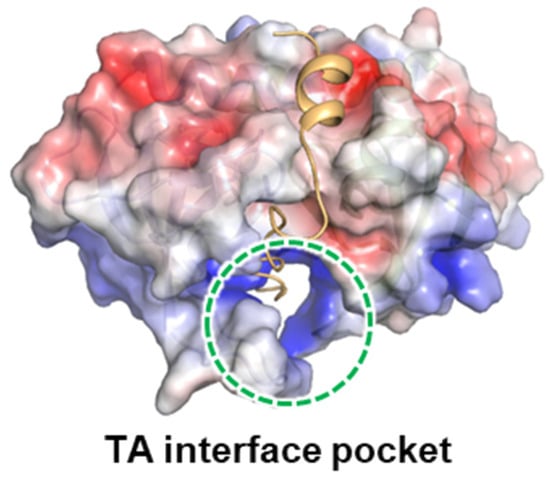
Figure 1
Open AccessArticle
Associations between Polycystic Ovary Syndrome (PCOS) and Antibiotic Use: Results from the UAEHFS
by
Nirmin F. Juber, Abdishakur Abdulle, Amar Ahmad, Fatme AlAnouti, Tom Loney, Youssef Idaghdour, Yvonne Valles and Raghib Ali
Antibiotics 2024, 13(5), 397; https://doi.org/10.3390/antibiotics13050397 - 26 Apr 2024
Abstract
Women with polycystic ovary syndrome (PCOS) have a higher susceptibility to infections compared to those without PCOS. Studies evaluating antibiotic use based on PCOS status are scarce. Therefore, we aimed to (i) assess the associations between self-reported PCOS and antibiotic use, and (ii)
[...] Read more.
Women with polycystic ovary syndrome (PCOS) have a higher susceptibility to infections compared to those without PCOS. Studies evaluating antibiotic use based on PCOS status are scarce. Therefore, we aimed to (i) assess the associations between self-reported PCOS and antibiotic use, and (ii) whether PCOS treatment and the age at PCOS diagnosis modified the associations above. This cross-sectional analysis used the United Arab Emirates Healthy Future Study (UAEHFS) conducted from February 2016 to March 2023 involving 2063 Emirati women aged 18–62 years. We performed ordinal logistic regressions under unadjusted and demographic-health-characteristic-adjusted models to obtain the odds ratios (ORs) and 95% confidence intervals (CIs) to analyze PCOS and antibiotic use. Subgroup analyses were performed by treatment status and age at diagnosis. We found that women with PCOS were 55% more likely to frequently take a course of antibiotics in the past year (aOR 1.55; 95% CI 1.26–1.90). Similar likelihoods were also found among those being treated for PCOS and those without treatment but with a PCOS diagnosis at ≤25 years. Our study suggests that PCOS was associated with an increased use of antibiotics among Emirati women. Understanding the frequent antibiotic use susceptibility among those with PCOS may improve antibiotic use surveillance and promote antibiotic stewardship in these at-risk individuals.
Full article
(This article belongs to the Special Issue Antimicrobial Prescribing, Population Use and Resistance, Impact in Global Health)
►▼
Show Figures
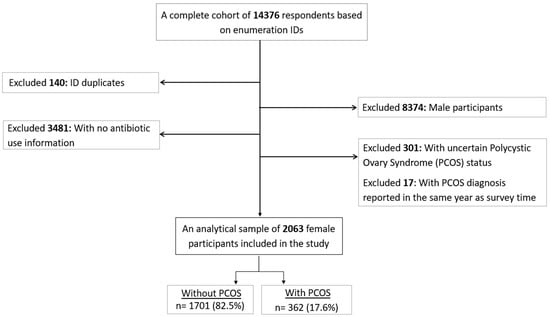
Figure 1
Open AccessArticle
Exploring the Molecular Mechanisms of Macrolide Resistance in Laboratory Mutant Helicobacter pylori
by
Meltem Ayaş, Sinem Oktem-Okullu, Orhan Özcan, Tanıl Kocagöz and Yeşim Gürol
Antibiotics 2024, 13(5), 396; https://doi.org/10.3390/antibiotics13050396 - 26 Apr 2024
Abstract
Resistance to clarithromycin, a macrolide antibiotic used in the first-line treatment of Helicobacter pylori infection, is the most important cause of treatment failure. Although most cases of clarithromycin resistance in H. pylori are associated with point mutations in 23S ribosomal RNA (rRNA), the
[...] Read more.
Resistance to clarithromycin, a macrolide antibiotic used in the first-line treatment of Helicobacter pylori infection, is the most important cause of treatment failure. Although most cases of clarithromycin resistance in H. pylori are associated with point mutations in 23S ribosomal RNA (rRNA), the relationships of other mutations with resistance remain unclear. We examined possible new macrolide resistance mechanisms in resistant strains using next-generation sequencing. Two resistant strains were obtained from clarithromycin-susceptible H. pylori following exposure to low clarithromycin concentrations using the agar dilution method. Sanger sequencing and whole-genome sequencing were performed to detect resistance-related mutations. Both strains carried the A2142G mutation in 23S rRNA. Candidate mutations (T1495A, T1494A, T1490A, T1476A, and G1472T) for clarithromycin resistance were detected in the Mutant-1 strain. Furthermore, a novel mutation in the gene encoding for the sulfite exporter TauE/SafE family protein was considered to be linked to clarithromycin resistance or cross-resistance, being identified as a target for further investigations. In the Mutant-2 strain, a novel mutation in the gene that encodes DUF874 family protein that can be considered as relevant with antibiotic resistance was detected. These mutations were revealed in the H. pylori genome for the first time, emphasizing their potential as targets for advanced studies.
Full article
(This article belongs to the Special Issue Helicobacter pylori Infection and Antibiotic Resistance: The Emerging Eradication Strategies)
►▼
Show Figures

Figure 1
Open AccessArticle
Low Oxygen Concentration Reduces Neisseria gonorrhoeae Susceptibility to Resazurin
by
Justin Rice, Jordan Gibson, Emily Young, Kendall Souder, Kailee Cunningham and Deanna M. Schmitt
Antibiotics 2024, 13(5), 395; https://doi.org/10.3390/antibiotics13050395 - 26 Apr 2024
Abstract
Neisseria gonorrhoeae has developed resistance to every antibiotic currently approved for the treatment of gonorrhea, prompting the development of new therapies. The phenoxazine dye resazurin exhibits robust antimicrobial activity against N. gonorrhoeae in vitro but fails to limit vaginal colonization by N. gonorrhoeae
[...] Read more.
Neisseria gonorrhoeae has developed resistance to every antibiotic currently approved for the treatment of gonorrhea, prompting the development of new therapies. The phenoxazine dye resazurin exhibits robust antimicrobial activity against N. gonorrhoeae in vitro but fails to limit vaginal colonization by N. gonorrhoeae in a mouse model. The lack of in vivo efficacy may be due to oxygen limitation as in vitro susceptibility assays with resazurin are conducted under atmospheric oxygen while a microaerophilic environment is present in the vagina. Here, we utilized broth microdilution assays to determine the susceptibility of N. gonorrhoeae to resazurin under low and atmospheric oxygen conditions. The minimal inhibitory concentration of resazurin for multiple N. gonorrhoeae clinical isolates was significantly higher under low oxygen. This effect was specific to resazurin as N. gonorrhoeae was equally susceptible to other antibiotics under low and atmospheric oxygen conditions. The reduced susceptibility of N. gonorrhoeae to resazurin under low oxygen was largely attributed to reduced oxidative stress, as the addition of antioxidants under atmospheric oxygen mimicked the reduced susceptibility to resazurin observed under low oxygen. Together, these data suggest oxygen concentration is an important factor to consider when evaluating the efficacy of new antibiotics against N. gonorrhoeae in vitro.
Full article
(This article belongs to the Special Issue Novel Approaches for the Treatment of Antimicrobial-Resistant Pathogens)
►▼
Show Figures
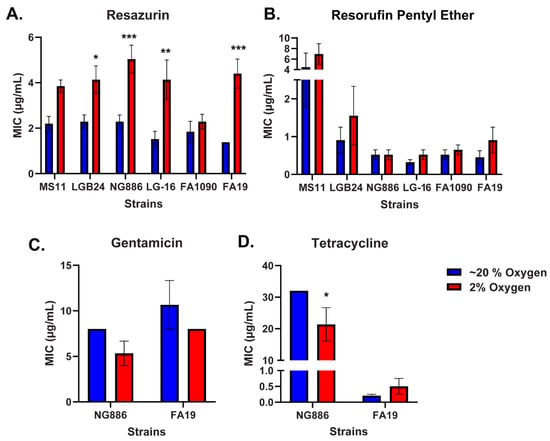
Figure 1
Open AccessCommunication
Detecting Class 1 Integrons and Their Variable Regions in Escherichia coli Whole-Genome Sequences Reported from Andean Community Countries
by
María Nicole Solis, Karen Loaiza, Lilibeth Torres-Elizalde, Ivan Mina, Miroslava Anna Šefcová and Marco Larrea-Álvarez
Antibiotics 2024, 13(5), 394; https://doi.org/10.3390/antibiotics13050394 (registering DOI) - 25 Apr 2024
Abstract
Various genetic elements, including integrons, are known to contribute to the development of antimicrobial resistance. Class 1 integrons have been identified in E. coli isolates and are associated with multidrug resistance in countries of the Andean Community. However, detailed information on the gene
[...] Read more.
Various genetic elements, including integrons, are known to contribute to the development of antimicrobial resistance. Class 1 integrons have been identified in E. coli isolates and are associated with multidrug resistance in countries of the Andean Community. However, detailed information on the gene cassettes located on the variable regions of integrons is lacking. Here, we investigated the presence and diversity of class 1 integrons, using an in silico approach, in 2533 whole-genome sequences obtained from EnteroBase. IntFinder v1.0 revealed that almost one-third of isolates contained these platforms. Integron-bearing isolates were associated with environmental, food, human, and animal origins reported from all countries under scrutiny. Moreover, they were identified in clones known for their pathogenicity or multidrug resistance. Integrons carried cassettes associated with aminoglycoside (aadA), trimethoprim (dfrA), cephalosporin (blaOXA; blaDHA), and fluoroquinolone (aac(6′)-Ib-cr; qnrB) resistance. These platforms showed higher diversity and larger numbers than previously reported. Moreover, integrons carrying more than three cassettes in their variable regions were determined. Monitoring the prevalence and diversity of genetic elements is necessary for recognizing emergent patterns of resistance in pathogenic bacteria, especially in countries where various factors are recognized to favor the selection of resistant microorganisms.
Full article
(This article belongs to the Special Issue Genomic Analysis of Antimicrobial Drug-Resistant Bacteria)
►▼
Show Figures
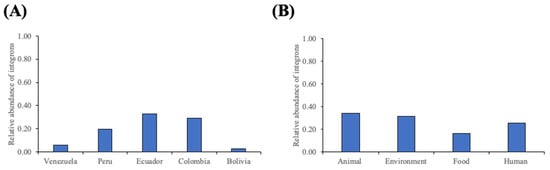
Figure 1
Open AccessArticle
New Insights into the Biological Functions of Essential TsaB/YeaZ Protein in Staphylococcus aureus
by
Haiyong Guo, Ting Lei, Junshu Yang, Yue Wang, Yifan Wang and Yinduo Ji
Antibiotics 2024, 13(5), 393; https://doi.org/10.3390/antibiotics13050393 (registering DOI) - 25 Apr 2024
Abstract
TsaB/YeaZ represents a promising target for novel antibacterial agents due to its indispensable role in bacterial survival, high conservation within bacterial species, and absence of eukaryotic homologs. Previous studies have elucidated the role of the essential staphylococcal protein, TsaB/YeaZ, in binding DNA to
[...] Read more.
TsaB/YeaZ represents a promising target for novel antibacterial agents due to its indispensable role in bacterial survival, high conservation within bacterial species, and absence of eukaryotic homologs. Previous studies have elucidated the role of the essential staphylococcal protein, TsaB/YeaZ, in binding DNA to mediate the transcription of the ilv-leu operon, responsible for encoding key enzymes involved in the biosynthesis of branched-chain amino acids—namely isoleucine, leucine, and valine (ILV). However, the regulation of ILV biosynthesis does not account for the essentiality of TsaB/YeaZ for bacterial growth. In this study, we investigated the impact of TsaB/YeaZ depletion on bacterial morphology and gene expression profiles using electron microscopy and deep transcriptomic analysis, respectively. Our results revealed significant alterations in bacterial size and surface smoothness upon TsaB/YeaZ depletion. Furthermore, we pinpointed specific genes and enriched biological pathways significantly affected by TsaB/YeaZ during the early and middle exponential phases and early stationary phases of growth. Crucially, our research uncovered a regulatory role for TsaB/YeaZ in bacterial autolysis. These discoveries offer fresh insights into the multifaceted biological functions of TsaB/YeaZ within S. aureus.
Full article
(This article belongs to the Special Issue Genomic Analysis of Antimicrobial Drug-Resistant Bacteria)
►▼
Show Figures
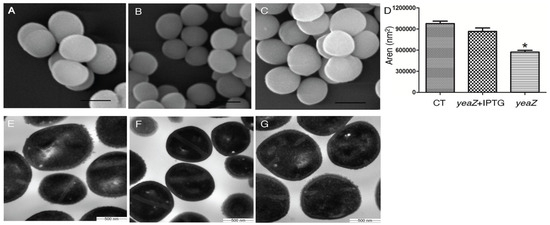

Figure 1
Open AccessReview
The Role of Gut Microbiota in the Etiopathogenesis of Multiple Chronic Diseases
by
Lara Pires, Ana M. González-Paramás, Sandrina A. Heleno and Ricardo C. Calhelha
Antibiotics 2024, 13(5), 392; https://doi.org/10.3390/antibiotics13050392 (registering DOI) - 25 Apr 2024
Abstract
►▼
Show Figures
Chronic diseases (CD) may result from a combination of genetic factors, lifestyle and social behaviours, healthcare system influences, community factors, and environmental determinants of health. These risk factors frequently coexist and interact with one another. Ongoing research and a focus on personalized interventions
[...] Read more.
Chronic diseases (CD) may result from a combination of genetic factors, lifestyle and social behaviours, healthcare system influences, community factors, and environmental determinants of health. These risk factors frequently coexist and interact with one another. Ongoing research and a focus on personalized interventions are pivotal strategies for preventing and managing chronic disease outcomes. A wealth of literature suggests the potential involvement of gut microbiota in influencing host metabolism, thereby impacting various risk factors associated with chronic diseases. Dysbiosis, the perturbation of the composition and activity of the gut microbiota, is crucial in the etiopathogenesis of multiple CD. Recent studies indicate that specific microorganism-derived metabolites, including trimethylamine N-oxide, lipopolysaccharide and uremic toxins, contribute to subclinical inflammatory processes implicated in CD. Various factors, including diet, lifestyle, and medications, can alter the taxonomic species or abundance of gut microbiota. Researchers are currently dedicating efforts to understanding how the natural progression of microbiome development in humans affects health outcomes. Simultaneously, there is a focus on enhancing the understanding of microbiome–host molecular interactions. These endeavours ultimately aim to devise practical approaches for rehabilitating dysregulated human microbial ecosystems, intending to restore health and prevent diseases. This review investigates how the gut microbiome contributes to CD and explains ways to modulate it for managing or preventing chronic conditions.
Full article
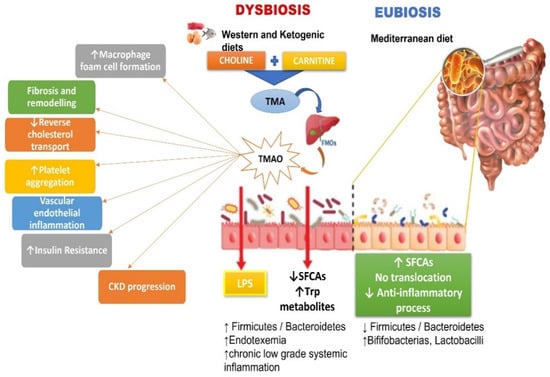
Figure 1
Open AccessReview
Extended-Spectrum Beta-Lactamase-Producing Escherichia coli and Other Antimicrobial-Resistant Gram-Negative Pathogens Isolated from Bovine Mastitis: A One Health Perspective
by
Breno Luis Nery Garcia, Stéfani Thais Alves Dantas, Kristian da Silva Barbosa, Thatiane Mendes Mitsunaga, Alyssa Butters, Carlos Henrique Camargo and Diego Borin Nobrega
Antibiotics 2024, 13(5), 391; https://doi.org/10.3390/antibiotics13050391 - 25 Apr 2024
Abstract
Antimicrobial resistance (AMR) poses an imminent threat to global public health, driven in part by the widespread use of antimicrobials in both humans and animals. Within the dairy cattle industry, Gram-negative coliforms such as Escherichia coli and Klebsiella pneumoniae stand out as major
[...] Read more.
Antimicrobial resistance (AMR) poses an imminent threat to global public health, driven in part by the widespread use of antimicrobials in both humans and animals. Within the dairy cattle industry, Gram-negative coliforms such as Escherichia coli and Klebsiella pneumoniae stand out as major causative agents of clinical mastitis. These same bacterial species are frequently associated with severe infections in humans, including bloodstream and urinary tract infections, and contribute significantly to the alarming surge in antimicrobial-resistant bacterial infections worldwide. Additionally, mastitis-causing coliforms often carry AMR genes akin to those found in hospital-acquired strains, notably the extended-spectrum beta-lactamase genes. This raises concerns regarding the potential transmission of resistant bacteria and AMR from mastitis cases in dairy cattle to humans. In this narrative review, we explore the distinctive characteristics of antimicrobial-resistant E. coli and Klebsiella spp. strains implicated in clinical mastitis and human infections. We focus on the molecular mechanisms underlying AMR in these bacterial populations and critically evaluate the potential for interspecies transmission. Despite some degree of similarity observed in sequence types and mobile genetic elements between strains found in humans and cows, the existing literature does not provide conclusive evidence to assert that coliforms responsible for mastitis in cows pose a direct threat to human health. Finally, we also scrutinize the existing literature, identifying gaps and limitations, and propose avenues for future research to address these pressing challenges comprehensively.
Full article
(This article belongs to the Special Issue Antimicrobial Resistance of Pathogens Isolated from Bovine Mastitis)
Open AccessArticle
Pseudomonas aeruginosa Infections in Patients with Severe COVID-19 in Intensive Care Units: A Retrospective Study
by
Alexandre Baudet, Marie Regad, Sébastien Gibot, Élodie Conrath, Julie Lizon, Béatrice Demoré and Arnaud Florentin
Antibiotics 2024, 13(5), 390; https://doi.org/10.3390/antibiotics13050390 - 25 Apr 2024
Abstract
Patients hospitalized in ICUs with severe COVID-19 are at risk for developing hospital-acquired infections, especially infections caused by Pseudomonas aeruginosa. We aimed to describe the evolution of P. aeruginosa infections in ICUs at CHRU-Nancy (France) in patients with severe COVID-19 during the
[...] Read more.
Patients hospitalized in ICUs with severe COVID-19 are at risk for developing hospital-acquired infections, especially infections caused by Pseudomonas aeruginosa. We aimed to describe the evolution of P. aeruginosa infections in ICUs at CHRU-Nancy (France) in patients with severe COVID-19 during the three initial waves of COVID-19. The second aims were to analyze P. aeruginosa resistance and to describe the antibiotic treatments. We conducted a retrospective cohort study among adult patients who were hospitalized for acute respiratory distress syndrome due to COVID-19 and who developed a hospital-acquired infection caused by P. aeruginosa during their ICU stay. Among the 51 patients included, most were male (90%) with comorbidities (77%), and the first identification of P. aeruginosa infection occurred after a median ICU stay of 11 days. Several patients acquired infections with MDR (27%) and XDR (8%) P. aeruginosa strains. The agents that strains most commonly exhibited resistance to were penicillin + β-lactamase inhibitors (59%), cephalosporins (42%), monobactams (32%), and carbapenems (27%). Probabilistic antibiotic treatment was prescribed for 49 patients (96%) and was subsequently adapted for 51% of patients after antibiogram and for 33% of patients after noncompliant antibiotic plasma concentration. Hospital-acquired infection is a common and life-threatening complication in critically ill patients. Efforts to minimize the occurrence and improve the treatment of such infections, including infections caused by resistant strains, must be pursued.
Full article
(This article belongs to the Special Issue Successful Antimicrobial Stewardship Approaches to Address Nosocomial Infections)
►▼
Show Figures
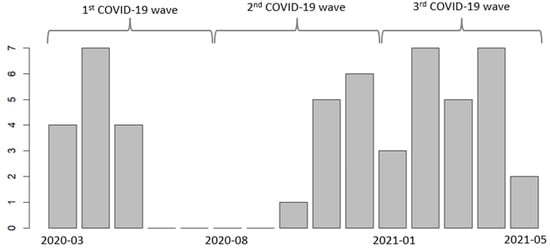
Figure 1

Journal Menu
► ▼ Journal Menu-
- Antibiotics Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Antibiotics, Antioxidants, JoF, Microbiology Research, Microorganisms
Redox in Microorganisms, 2nd Edition
Topic Editors: Michal Letek, Volker BehrendsDeadline: 31 July 2024
Topic in
Antibiotics, JPM, Pharmaceuticals, Pharmaceutics
Pharmacokinetic and Pharmacodynamic Modelling in Drug Discovery and Development
Topic Editors: Inaki F. Troconiz, Victor Mangas Sanjuán, Maria Garcia-Cremades MiraDeadline: 31 August 2024
Topic in
Molecules, Pharmaceutics, Antibiotics, Microorganisms, Biomolecules, Marine Drugs, Polymers, IJMS
Antimicrobial Agents and Nanomaterials
Topic Editors: Sandra Pinto, Vasco D. B. BonifácioDeadline: 30 September 2024
Topic in
Antibiotics, Biomedicines, JCM, Pharmaceuticals, Pharmaceutics
Challenges and Future Prospects of Antibacterial Therapy
Topic Editors: Kwang-sun Kim, Zehra EdisDeadline: 31 October 2024

Conferences
Special Issues
Special Issue in
Antibiotics
When Communities Matter: Interplay between Mobile Genetic Elements and Antibiotic Resistance
Guest Editor: Antony T. VincentDeadline: 30 April 2024
Special Issue in
Antibiotics
Antibacterial Treatment in Periodontal and Endodontic Therapy, 2nd Volume
Guest Editors: Andreas Braun, Felix KrauseDeadline: 15 May 2024
Special Issue in
Antibiotics
The Application of Antibiotic Therapy in Oral Surgery and Dental Implant Procedures
Guest Editors: Eugenio Velasco-Ortega, Angel-Orion Salgado-Peralvo, Juan Francisco Peña-CardellesDeadline: 31 May 2024
Special Issue in
Antibiotics
Challenges for Therapeutic Drug Monitoring of Antimicrobials
Guest Editors: Giuseppe Pipitone, Giuseppe Nunnari, Fabrizio TagliettiDeadline: 15 June 2024
Topical Collections
Topical Collection in
Antibiotics
Antimicrobial Resistance and Anti-Biofilms
Collection Editors: Ding-Qiang Chen, Yulong Tan, Ren-You Gan, Guanggang Qu, Zhenbo Xu, Junyan Liu








