Journal Description
Plants
Plants
is an international, scientific, peer-reviewed, open access journal on plant science published semimonthly online by MDPI. The Australian Society of Plant Scientists (ASPS), the Spanish Phytopathological Society (SEF), the Spanish Society of Plant Physiology (SEFV), the Spanish Society of Horticultural Sciences (SECH) and the Italian Society of Phytotherapy (S.I.Fit.) are affiliated with Plants and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, PubAg, AGRIS, CAPlus / SciFinder, and other databases.
- Journal Rank: JCR - Q1 (Plant Sciences) / CiteScore - Q1 (Plant Science)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 15.3 days after submission; acceptance to publication is undertaken in 2.8 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
4.5 (2022);
5-Year Impact Factor:
4.8 (2022)
Latest Articles
Chitosan-GSNO Nanoparticles and Silicon Priming Enhance the Germination and Seedling Growth of Soybean (Glycine max L.)
Plants 2024, 13(10), 1290; https://doi.org/10.3390/plants13101290 (registering DOI) - 07 May 2024
Abstract
Soybean, a major legume crop, has seen a decline in its production owing to challenges in seed germination and the development of seedlings. Thus, in this study, we systematically investigated the influence of various chitosan–S-nitrosoglutathione (chitosan-GSNO) nanoparticle (0, 25, 50, and 100 µM)
[...] Read more.
Soybean, a major legume crop, has seen a decline in its production owing to challenges in seed germination and the development of seedlings. Thus, in this study, we systematically investigated the influence of various chitosan–S-nitrosoglutathione (chitosan-GSNO) nanoparticle (0, 25, 50, and 100 µM) and Si (0, 0.5, and 1 mM) priming concentrations on soybean seed germination and seedling growth over five different priming durations (range: 1–5 h at each concentration). Significant differences were observed in all parameters, except seedling diameter, with both treatments. Seed germination was significantly enhanced after 3 h of priming in both treatments. The final germination percentage (FGP), peak germination percentage (PGP), vigor index (VI), seedling biomass (SB), hypocotyl length (HL), and radical length (RL) of 100 μM chitosan-GSNO-nanoparticle-primed seeds increased by 20.3%, 41.3%, 78.9%, 25.2%, 15.7%, and 65.9%, respectively, compared with those of the control; however, the mean germination time (MGT) decreased by 18.43%. Si priming at 0.5 mM increased the FGP, PGP, VI, SB, HL, and RL by 13.9%, 55.17%, 39.2%, 6.5%, 22.5%, and 25.1%, respectively, but reduced the MGT by 12.29% compared with the control treatment. Chitosan-GSNO and Si treatment up-regulated the relative expression of gibberellic acid (GA)-related genes (GmGA3ox3 and GmGA2ox1) and down-regulated that of abscisic acid (ABA)-related genes (GmABA2, GmAAO3, and GmNCED5). Chitosan-GSNO and Si application increased bioactive GA4 levels and simultaneously reduced ABA content. Hence, the use of exogenous chitosan-GSNO nanoparticles and Si as priming agents had a beneficial effect on seed germination and seedling growth because of the up-regulation in the expression of GA and down-regulation in the expression of ABA. Additional research is needed to understand the combined impact of Si and chitosan-GSNO nanoparticles, including their effects on the expression levels of other hormones and genes even in the later growth stage of the crop.
Full article
(This article belongs to the Special Issue Studies on Strategy of Diaspore Dispersal and Effectiveness of Seed Germination)
►
Show Figures
Open AccessArticle
A 14-3-3 Protein Ca16R Acts Positively in Pepper Immunity against Ralstonia solanacearum by Interacting with CaASR1
by
Sheng Yang, Meiyun Wan, Xingge Cheng, Qing Cheng and Huolin Shen
Plants 2024, 13(10), 1289; https://doi.org/10.3390/plants13101289 - 07 May 2024
Abstract
Although 14-3-3 proteins have been implicated in plant growth, development, and stress response, their roles in pepper immunity against R. solanacearum remain poorly understood. In this study, a 14-3-3-encoding gene in pepper, Ca16R, was found to be upregulated by R. solanacearum inoculation
[...] Read more.
Although 14-3-3 proteins have been implicated in plant growth, development, and stress response, their roles in pepper immunity against R. solanacearum remain poorly understood. In this study, a 14-3-3-encoding gene in pepper, Ca16R, was found to be upregulated by R. solanacearum inoculation (RSI), its silencing significantly reduced the resistance of pepper plants to RSI, and its overexpression significantly enhanced the resistance of Nicotiana benthamiana to RSI. Consistently, its transient overexpression in pepper leaves triggered HR cell death, indicating that it acts positively in pepper immunity against RSI, and it was further found to act positively in pepper immunity against RSI by promoting SA but repressing JA signaling. Ca16R was also found to interact with CaASR1, originally using pull-down combined with a spectrum assay, and then confirmed using bimolecular fluorescence complementation (BiFC) and a pull-down assay. Furthermore, we found that CaASR1 transient overexpression induced HR cell death and SA-dependent immunity while repressing JA signaling, although this induction and repression was blocked by Ca16R silencing. All these data indicate that Ca16R acts positively in pepper immunity against RSI by interacting with CaASR1, thereby promoting SA-mediated immunity while repressing JA signaling. These results provide new insight into mechanisms underlying pepper immunity against RSI.
Full article
(This article belongs to the Special Issue Genetics of Disease Resistance in Horticultural Crops)
►▼
Show Figures

Figure 1
Open AccessArticle
Production of Polyphenolic Natural Products by Bract-Derived Tissue Cultures of Three Medicinal Tilia spp.: A Comparative Untargeted Metabolomics Study
by
Zsolt Szűcs, Zoltán Cziáky, László Volánszki, Csaba Máthé, Gábor Vasas and Sándor Gonda
Plants 2024, 13(10), 1288; https://doi.org/10.3390/plants13101288 - 07 May 2024
Abstract
Medicinal plant tissue cultures are potential sources of bioactive compounds. In this study, we report the chemical characterization of the callus cultures of three medicinal Tilia spp. (Tilia cordata, Tilia vulgaris and Tilia tomentosa), along with the comparison to bracts
[...] Read more.
Medicinal plant tissue cultures are potential sources of bioactive compounds. In this study, we report the chemical characterization of the callus cultures of three medicinal Tilia spp. (Tilia cordata, Tilia vulgaris and Tilia tomentosa), along with the comparison to bracts and flowers of the same species. Our aim was to show that calli of Tilia spp. are good alternatives to the calli of T. americana for the production of polyphenols and are better sources of a subset of polyphenolic metabolites, compared to the original organs. Calli were initiated from young bracts and grown on woody plant medium containing 1 mg L−1 2,4-D and 0.1 mg L−1 BAP. For chemical characterization, a quality-controlled untargeted metabolomics approach and the quantification of several bioactive compounds was performed with the use of LC-ESI-MS/MS. While bracts and flowers contained flavonoid glycosides (astragalin, isoquercitrin) as major polyphenols, calli of all species contained catechins, coumarins (fraxin, esculin and scopoletin) and flavane aglyca. T. tomentosa calli contained 5397 µg g DW−1 catechin, 201 µg g DW−1 esculin, 218 µg g DW−1 taxifolin and 273 µg g DW−1 eriodictyol, while calli from other species contained lower amounts. T. cordata and T. tomentosa flowers were rich in isoquercitrin, containing 8134 and 6385 µg g DW−1, respectively. The currently tested species contained many of the bioactive metabolites described from T. americana. The production of catechin was shown to be comparable to the most efficient tissue cultures reported. Flowers and bracts contained flavonoid glycosides, including tiliroside, resembling bioactive fractions of T. americana. In addition, untargeted metabolomics has shown fingerprint-like differences among species, highlighting possible chemotaxonomic and quality control applications, especially for bracts.
Full article
(This article belongs to the Special Issue Plant Tissue Culture and Secondary Metabolites)
Open AccessArticle
Transcriptome Analysis Revealed the Possible Reasons for the Change of Ni Resistance in Rhus typhina after Spraying Melatonin
by
Tongbao Qu, Yinxi Ma, Minqiang Yun and Chunli Zhao
Plants 2024, 13(10), 1287; https://doi.org/10.3390/plants13101287 - 07 May 2024
Abstract
Melatonin (MT) plays an important role in alleviating the stress of soil heavy metal pollution on plants. However, its ability to improve the tolerance of Rhus typhina to Ni stress and its mechanism of action are still unclear. Therefore, MT (0, 50, 100,
[...] Read more.
Melatonin (MT) plays an important role in alleviating the stress of soil heavy metal pollution on plants. However, its ability to improve the tolerance of Rhus typhina to Ni stress and its mechanism of action are still unclear. Therefore, MT (0, 50, 100, and 200 μmol·L−1) was sprayed on the leaf surface of R. typhina seedlings under Ni (0 and 250 mg·kg−1) stress to study the differences in growth, physiology, and gene expression. The results showed that exogenous MT could improve the ability of R. typhina to resist Ni stress by inhibiting the degradation of chlorophyll and carotenoid, enhancing photosynthesis, and augmenting the activity of antioxidant enzymes. Moreover, 100 μmol·L−1 MT could increase the Ni concentration in R. typhina seedlings and reduce the translocation factor. Transcriptome analysis showed that MT mainly regulated the expression of related genes in plant hormone signal transduction, starch and sucrose metabolism, and various amino acid metabolism pathways. This study combined physiological and transcriptomic analysis to reveal the molecular mechanism of MT enhancing Ni resistance in R. typhina, and provides a new direction for expanding its application in phytoremediation.
Full article
(This article belongs to the Special Issue Adaptive Strategies of Plants to Stress Factors)
Open AccessArticle
Genome-Wide Analysis of the SAUR Gene Family and Its Expression Profiles in Response to Salt Stress in Santalum album
by
Qing Zhu, Haoyue Zheng, Xu Hu, Yi Liu, Xinyi Zheng, Libei Li and Minqiang Tang
Plants 2024, 13(10), 1286; https://doi.org/10.3390/plants13101286 - 07 May 2024
Abstract
The SAUR (small auxin-up RNA) family constitutes a category of genes that promptly respond to the hormone auxin and play a pivotal role in diverse biological processes encompassing plant growth and the response to abiotic stress. Santalum album L., a semi-parasitic evergreen tree,
[...] Read more.
The SAUR (small auxin-up RNA) family constitutes a category of genes that promptly respond to the hormone auxin and play a pivotal role in diverse biological processes encompassing plant growth and the response to abiotic stress. Santalum album L., a semi-parasitic evergreen tree, is renowned for its economically valuable essential oils, positioning it among the most prized tree species. In this study, a meticulous identification and comprehensive analysis of 43 SAUR genes was conducted within S. album. Based on phylogenetic relationships, the SaSAUR genes were systematically categorized into five groups. A collinearity analysis revealed intriguing insights, disclosing 14 segmental duplications and 9 tandem duplications within the SaSAUR genes, emphasizing the pivotal role of duplication in the expansion of this gene family. Noteworthy variations in the expression levels of SaSAUR genes were observed by delving into the SaSAUR transcriptome data from various tissues, including leaves, roots, and heartwood, as well as under salt-stress conditions. Notably, SaSAUR08 and SaSAUR13 were significantly upregulated in heartwood compared with roots and leaves, while SaSAUR18 was markedly more expressed in roots compared with heartwood and leaves. Furthermore, SaSAUR27 and SaSAUR28 were found to respond closely to salt stress, hinting at their potential involvement in the salt-stress response mechanism. This research offers a comprehensive investigation of SAUR genes in S. album and establishes a foundation for future exploration of the SAUR gene family, particularly its relation to growth and salt-stress responses.
Full article
(This article belongs to the Special Issue Crop Functional Genomics and Biological Breeding)
►▼
Show Figures

Figure 1
Open AccessArticle
QTL Mapping of Yield-Related Traits in Tetraploid Wheat Based on Wheat55K SNP Array
by
Yatao Jia, Yifan Zhang, Yingkai Sun, Chao Ma, Yixiong Bai, Hanbing Zhang, Junbin Hou, Yong Wang, Wanquan Ji, Haibo Bai, Shuiyuan Hao and Zhonghua Wang
Plants 2024, 13(10), 1285; https://doi.org/10.3390/plants13101285 - 07 May 2024
Abstract
To enhance the understanding of yield-related traits in tetraploid wheat, it is crucial to investigate and identify genes that govern superior yield characteristics. This study utilized the wheat55K single nucleotide polymorphism array to genotype a recombinant inbred line (RIL) population consisting of 120
[...] Read more.
To enhance the understanding of yield-related traits in tetraploid wheat, it is crucial to investigate and identify genes that govern superior yield characteristics. This study utilized the wheat55K single nucleotide polymorphism array to genotype a recombinant inbred line (RIL) population consisting of 120 lines developed through the crossbreeding of two tetraploid wheat varieties, Qin Hei-1 (QH-1) and Durum Wheat (DW). An investigation and analysis were conducted on 11 yield-related traits, including peduncle length (PL), neck length (NL), spike length (SL), flowering date (FD), heading date (HD), thousand-kernel weight (TKW), kernel area ratio (KAR), kernel circumference (KC), kernel length (KL), kernel width (KW), and kernel length–width ratio (KL-WR), over a period of three years in two locations, Yang Ling, Shaanxi, and Lin He, Inner Mongolia. The analysis identified nine stable loci among eight agronomic traits, named QSL.QD-1A.1, QNL.QD-4B.2, QPL.QD-4B.1, QFD.QD-2B, QHD.QD-2B.1, QHD.QD-4B, QKC.QD-4B.2, QKL-WR.QD-4B.6, and QKL.QD-4B.2. Among them, the additive effects of three QTLs, QSL.QD-1A.1, QNL.QD-4B.2, and QFD.QD-2B, were positive, indicating that the enhancing alleles at these loci were derived from the parent line QH-1. These three QTLs showed significant positive effects on the phenotypes of the population materials. Furthermore, potential functional genes were identified within the mapping intervals of QSL.QD-1A.1 and QNL.QD-4B.2, which regulate the development of spike length and neck length, respectively. These results provide potential QTLs and candidate genes, which broaden the genetic basis of agronomic traits related to yield, such as SL, NL, PL, and FD, and benefits for wheat breeding and improvement.
Full article
(This article belongs to the Special Issue Genetics, Genomics, and Biotechnology for Cereal Crop Improvements)
Open AccessArticle
Molecular Mechanisms of Chlorophyll Deficiency in Ilex × attenuata ‘Sunny Foster’ Mutant
by
Yiping Zou, Yajian Huang, Donglin Zhang, Hong Chen, Youwang Liang, Mingzhuo Hao and Yunlong Yin
Plants 2024, 13(10), 1284; https://doi.org/10.3390/plants13101284 - 07 May 2024
Abstract
Ilex × attenuata ‘Sunny Foster’ represents a yellow leaf mutant originating from I. × attenuata ‘Foster#2’, a popular ornamental woody cultivar. However, the molecular mechanisms underlying this leaf color mutation remain unclear. Using a comprehensive approach encompassing cytological, physiological, and transcriptomic methodologies, notable
[...] Read more.
Ilex × attenuata ‘Sunny Foster’ represents a yellow leaf mutant originating from I. × attenuata ‘Foster#2’, a popular ornamental woody cultivar. However, the molecular mechanisms underlying this leaf color mutation remain unclear. Using a comprehensive approach encompassing cytological, physiological, and transcriptomic methodologies, notable distinctions were discerned between the mutant specimen and its wild type. The mutant phenotype displayed aberrant chloroplast morphology, diminished chlorophyll content, heightened carotenoid/chlorophyll ratios, and a decelerated rate of plant development. Transcriptome analysis identified differentially expressed genes (DEGs) related to chlorophyll metabolism, carotenoid biosynthesis and photosynthesis. The up-regulation of CHLD and CHLI subunits leads to decreased magnesium chelatase activity, while the up-regulation of COX10 increases heme biosynthesis—both impair chlorophyll synthesis. Conversely, the down-regulation of HEMD hindered chlorophyll synthesis, and the up-regulation of SGR enhanced chlorophyll degradation, resulting in reduced chlorophyll content. Additionally, genes linked to carotenoid biosynthesis, flavonoid metabolism, and photosynthesis were significantly down-regulated. We also identified 311 putative differentially expressed transcription factors, including bHLHs and GLKs. These findings shed light on the molecular mechanisms underlying leaf color mutation in I. × attenuata ‘Sunny Foster’ and provide a substantial gene reservoir for enhancing leaf color through breeding techniques.
Full article
(This article belongs to the Section Horticultural Science and Ornamental Plants)
Open AccessArticle
Quantitative Determination of Biogenic Element Contents and Phytochemicals of Broccoli (Brassica oleracea var. italica) Cooked Using Different Techniques
by
Fahad AlJuhaimi, Isam A. Mohamed Ahmed, Mehmet Musa Özcan, Nurhan Uslu and Zainab Albakry
Plants 2024, 13(10), 1283; https://doi.org/10.3390/plants13101283 - 07 May 2024
Abstract
In this study, the effect of different cooking techniques on broccoli moisture, total phenolic, total flavonoid, and radical scavenging capacity results, polyphenol contents, and their quantitative values was investigated. The total phenolic quantities of fresh and cooked broccoli samples were assessed to be
[...] Read more.
In this study, the effect of different cooking techniques on broccoli moisture, total phenolic, total flavonoid, and radical scavenging capacity results, polyphenol contents, and their quantitative values was investigated. The total phenolic quantities of fresh and cooked broccoli samples were assessed to be between 36.32 (conventional boiling) and 423.39 mg GAE/100 g (microwave heating). The radical scavenging activities of the broccoli samples were reported between 2.55 (conventional boiling) and 4.99 mmol/kg (microwave heating). In addition, catechin and rutin quantities of the fresh and cooked broccoli samples were measured to be between 2.24 (conventional boiling) and 54.48 mg/100 g (microwave heating), and between 0.55 (conventional boiling) and 16.33 mg/100 g (microwave heating), respectively. The most abundant elements in fresh and cooked broccoli samples were K, Ca, P, S, and Mg. The results showed some changes depending on cooking techniques compared to the control. The bioactive properties of broccoli samples cooked by means of conventional boiling, boiling in vacuum bag, and high-pressure boiling were established to be lower compared to the fresh sample. Catechin, 3,4-dihydroxybenzoic acid, rutin, and gallic acid were the key phenolic compounds of fresh and cooked broccoli samples. The phenolic components of broccoli were significantly affected by the applied cooking techniques. The highest protein in broccoli samples was determined in the broccoli sample cooked by boiling in a vacuum bag. There were statistically significant changes among the mineral results of broccoli cooked with different cooking methods.
Full article
(This article belongs to the Special Issue Phytochemicals in Plants: Recent Developments on the Occurrence, Composition, Stability, Health, Food and Pharmaceutical Applications—2nd Edition)
Open AccessArticle
Relicts of Threatened Biodiversity: Similarities and Differences among the 7230 EU Habitat Plant Communities on Montane Plateaus of Central Apennines, Italy
by
Giampiero Ciaschetti, Safiya Praleskouskaya and Roberto Venanzoni
Plants 2024, 13(10), 1282; https://doi.org/10.3390/plants13101282 - 07 May 2024
Abstract
The habitats protected by the European Union (EU) include most peat vegetation, such as mires, swamp mires, fens, and peat bogs—all belonging to the classes Oxycocco–Sphagnetea and Scheuchzerio–Caricetea fuscae and carrying the Habitat Codes 71xx and 72xx. These types of
[...] Read more.
The habitats protected by the European Union (EU) include most peat vegetation, such as mires, swamp mires, fens, and peat bogs—all belonging to the classes Oxycocco–Sphagnetea and Scheuchzerio–Caricetea fuscae and carrying the Habitat Codes 71xx and 72xx. These types of vegetation are typical of cold and cool temperate climates, while they become rarer in Southern Europe where Mediterranean influences prevail, representing relic fragments of the past glacial climatic conditions there. Because of their limited extension and the increasing warmth and drought due to climate change, they are seriously threatened. Even if many studies were performed, their richness and distribution across Europe are still not well–understood, and only a few examples are known from the Central and Southern Apennines to date. In order to provide the syntaxonomical classification of the alkaline fens referable to the EU Habitat 7230 found on the mountain plateaus of the Central Apennines, we analyzed their species structure and flora composition, together with their chorological and ecological characteristics. We also evaluated their conservation status, pressures, and threats. The alkaline fens of the Central Apennines are found to be poorer in diagnostic species when compared to similar communities of Central and Northern Europe. However, they are rich in the species of the surrounding meadows and pastures. Among them, the new subassociation Caricetum davallianae caricetosum hostianae is described.
Full article
(This article belongs to the Special Issue Advanced Botanical Research in the Mediterranean Area: Studies in Honor of Prof. Francesco Maria Raimondo on the Occasion of His 80th Birthday)
►▼
Show Figures

Figure 1
Open AccessArticle
New Understanding of Meta-Topolin Riboside Metabolism in Micropropagated Woody Plants
by
Maroua Grira, Els Prinsen and Stefaan Werbrouck
Plants 2024, 13(9), 1281; https://doi.org/10.3390/plants13091281 - 06 May 2024
Abstract
Topolin cytokinins have emerged as valuable tools in micropropagation. This study investigates the metabolism of meta-topolin riboside (mTR) in three distinct tree species: Handroanthus guayacan and Tabebuia rosea (Bignoniaceae), and Tectona grandis (Lamiaceae). Employing labeled N15 mTR, we
[...] Read more.
Topolin cytokinins have emerged as valuable tools in micropropagation. This study investigates the metabolism of meta-topolin riboside (mTR) in three distinct tree species: Handroanthus guayacan and Tabebuia rosea (Bignoniaceae), and Tectona grandis (Lamiaceae). Employing labeled N15 mTR, we unraveled the complex mechanisms underlying cytokinin homeostasis, identifying N9-glucosylation as the principal deactivation pathway. Our findings demonstrate a capacity in T. rosea and H. guayacan to reposition the hydroxyl group on the cytokinin molecule, a previously unexplored metabolic pathway. Notably, this study reveals remarkable interfamilial and interspecies differences in mTR metabolism, challenging established perspectives on the role of callus tissue in cytokinin storage. These insights not only illuminate the metabolic intricacies of mTR, a cytokinin with interesting applications in plant tissue culture, but also enhances our understanding of cytokinin dynamics in plant systems, thereby enriching the scientific discourse on plant physiology and cytokinin biology.
Full article
(This article belongs to the Special Issue Plant Propagation)
►▼
Show Figures

Figure 1
Open AccessTechnical Note
The Effect of Temperature on the Inflorescence Formation Model for Phalaenopsis
by
Jiunyuan Chen and Chiachung Chen
Plants 2024, 13(9), 1280; https://doi.org/10.3390/plants13091280 - 06 May 2024
Abstract
Phalaenopsis orchids are a popular ornamental plant in the flower market. During some festivals, demand increases significantly. These mature orchids must be placed in cooling rooms for inflorescence formation at specific times to increase the financial return from their sale. The purpose of
[...] Read more.
Phalaenopsis orchids are a popular ornamental plant in the flower market. During some festivals, demand increases significantly. These mature orchids must be placed in cooling rooms for inflorescence formation at specific times to increase the financial return from their sale. The purpose of this study is to evaluate the effect of day and night temperatures on the inflorescence formation percentage using the proposed sigmoid model. Four varieties that are cultured in different vegetative temperature regimes are placed in a cooling room. An empirical inflorescence formation model is proposed as a management tool to predict the inflorescence formation percentage for Phalaenopsis. Some data sets from previous studies are used for comparison. The accumulation temperature is calculated using the day and night temperatures and is an index to predict the inflorescence formation percentage. The results show that there is a similar distribution of the inflorescence formation percentage and accumulation temperature for the four varieties. The proposed sigmoid model has a good fitting ability for the inflorescence formation percentage. This inflorescence formation model from the pooled data sets allows quantitative microclimate management of the vegetative and cooling room.
Full article
(This article belongs to the Special Issue Modelling for Prediction of Horticultural Plant Growth and Defense)
►▼
Show Figures

Figure 1
Open AccessArticle
Microplastic Has No Effect on Rice Yield and Gaseous N Emission from an Infertile Soil with High Inorganic N Inputs
by
Si Wu, Haiying Lu, Zhenghua Yi, Gui Chen and Haijun Sun
Plants 2024, 13(9), 1279; https://doi.org/10.3390/plants13091279 - 06 May 2024
Abstract
Microplastic might affect the crop yield, nitrogen (N) use efficiency and reactive N losses from agricultural soil systems. However, evaluation of these effects in infertile soil planted with different rice cultivars is lacking. We conducted a soil column experiment to determine the influence
[...] Read more.
Microplastic might affect the crop yield, nitrogen (N) use efficiency and reactive N losses from agricultural soil systems. However, evaluation of these effects in infertile soil planted with different rice cultivars is lacking. We conducted a soil column experiment to determine the influence of a typical microplastic polyethylene (PE) input into an infertile soil with 270 kg N ha−1 and planted with two rice cultivars, i.e., a common rice Nangeng 5055 (NG) and a hybrid rice Jiafengyou 6 (JFY). The results showed that JFY produced a significantly (p < 0.05) greater grain yield than NG (61.6–66.2 vs. 48.2–52.5 g pot−1) but was not influenced by PE. Overall, PE hardly changed the N use efficiency of NG and JFY. Unexpectedly, PE significantly (p < 0.05) increased the total amino acid content of NG. Compared with JFY, NG volatilized significantly (p < 0.05) more ammonia (NH3) (0.84–0.92 vs. 0.64–0.67 g N pot−1) but emitted equal nitrous oxide (N2O). PE exerted no effect on either NH3 volatilization or the N2O emission flux pattern and cumulative losses of the rice growth cycle, whether with NG or JFY. Some properties of tested soils changed after planting with different rice cultivars and incorporating with microplastic. In conclusion, the rice production, N use efficiency, NH3 volatilization and N2O emission from the N-fertilized infertile soil were pronouncedly influenced by the rice cultivar, but not the PE. However, PE influenced the grain quality of common rice and some properties of tested soils with both rice cultivars.
Full article
(This article belongs to the Section Plant–Soil Interactions)
►▼
Show Figures

Figure 1
Open AccessArticle
Micropropagation and Genetic Fidelity of Fegra Fig (Ficus palmata Forssk.) and Grafting Compatibility of the Regenerated Plants with Ficus carica
by
Ahmed Ali Al-Aizari, Yaser Hassan Dewir, Abdel-Halim Ghazy, Abdullah Al-Doss and Rashid Sultan Al-Obeed
Plants 2024, 13(9), 1278; https://doi.org/10.3390/plants13091278 - 06 May 2024
Abstract
Ficus palmata is an important fig species that produces edible and nutritious fruit and possesses several therapeutic uses. This study reports an effective method for the micropropagation of F. palmata using nodal explants. In vitro shoots were cultured for 7 weeks onto MS
[...] Read more.
Ficus palmata is an important fig species that produces edible and nutritious fruit and possesses several therapeutic uses. This study reports an effective method for the micropropagation of F. palmata using nodal explants. In vitro shoots were cultured for 7 weeks onto MS medium fortified with different concentrations of cytokinins, light intensities, sucrose concentrations, and light/dark incubation treatments. Optimal axillary shoot proliferation (10.9 shoots per explant) was obtained on a medium containing 30 g/L sucrose and supplemented with 2 mg/L 6-benzylaminopurine (BAP) under 35 μmol/m2/s light intensity. Dark incubation limited the foliage growth but favored shoot elongation and rooting compared with light incubation. Elongated shoots, under dark conditions, were rooted (100%; 6.67 roots per explant) onto MS medium containing 1 mg/L indole-3-acetic acid (IAA) and 1.5 g/L activated charcoal. The micropropagated plantlets were acclimatized with a 95% survival rate. In this study, the genetic fidelity of micropropagated F. palmata clones along with their mother plant was tested using randomly amplified polymorphic DNA (RAPD), inter-simple sequence repeats (ISSR), and start codon targeted (SCoT) molecular markers. The genetic similarity between the micropropagated plantlets and the mother plant of F. palmata was nearly 95.9%, assuring high uniformity and true-to-type regenerated plants. Using micropropagated F. palmata plantlets as a rootstock proved appropriate for the grafting F. carica ‘Brown Turkey’. These findings contribute to the commercial propagation and production of the fig crop.
Full article
(This article belongs to the Section Plant Genetics, Genomics and Biotechnology)
►▼
Show Figures
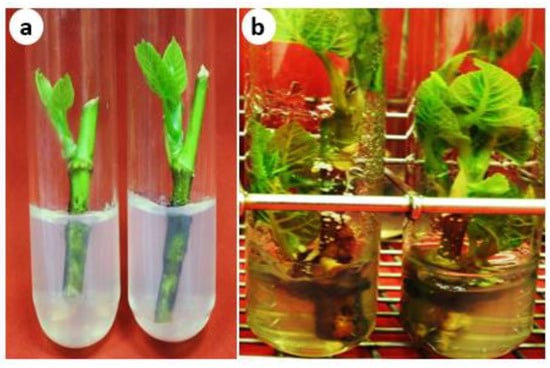

Figure 1
Open AccessArticle
Gibberellin Positively Regulates Tomato Resistance to Tomato Yellow Leaf Curl Virus (TYLCV)
by
Chenwei Zhang, Dandan Wang, Yan Li, Zifan Wang, Zhiming Wu, Qingyin Zhang, Hongwei Jia, Xiaoxu Dong, Lianfen Qi, Jianhua Shi and Zhonglin Shang
Plants 2024, 13(9), 1277; https://doi.org/10.3390/plants13091277 - 06 May 2024
Abstract
Tomato yellow leaf curl virus (TYLCV) is a prominent viral pathogen that adversely affects tomato plants. Effective strategies for mitigating the impact of TYLCV include isolating tomato plants from the whitefly, which is the vector of the virus, and utilizing transgenic lines that
[...] Read more.
Tomato yellow leaf curl virus (TYLCV) is a prominent viral pathogen that adversely affects tomato plants. Effective strategies for mitigating the impact of TYLCV include isolating tomato plants from the whitefly, which is the vector of the virus, and utilizing transgenic lines that are resistant to the virus. In our preliminary investigations, we observed that the use of growth retardants increased the rate of TYLCV infection and intensified the damage to the tomato plants, suggesting a potential involvement of gibberellic acid (GA) in the conferring of resistance to TYLCV. In this study, we employed an infectious clone of TYLCV to inoculate tomato plants, which resulted in leaf curling and growth inhibition. Remarkably, this inoculation also led to the accumulation of GA3 and several other phytohormones. Subsequent treatment with GA3 effectively alleviated the TYLCV-induced leaf curling and growth inhibition, reduced TYLCV abundance in the leaves, enhanced the activity of antioxidant enzymes, and lowered the reactive oxygen species (ROS) levels in the leaves. Conversely, the treatment with PP333 exacerbated TYLCV-induced leaf curling and growth suppression, increased TYLCV abundance, decreased antioxidant enzyme activity, and elevated ROS levels in the leaves. The analysis of the gene expression profiles revealed that GA3 up-regulated the genes associated with disease resistance, such as WRKYs, NACs, MYBs, Cyt P450s, and ERFs, while it down-regulated the DELLA protein, a key agent in GA signaling. In contrast, PP333 induced gene expression changes that were the opposite of those caused by the GA3 treatment. These findings suggest that GA plays an essential role in the tomato’s defense response against TYLCV and acts as a positive regulator of ROS scavenging and the expression of resistance-related genes.
Full article
(This article belongs to the Section Horticultural Science and Ornamental Plants)
►▼
Show Figures

Figure 1
Open AccessArticle
Identification of Novel Regulators of Leaf Senescence Using a Deep Learning Model
by
Chaocheng Guo, Zhuoran Huang, Jiahao Chen, Guolong Yu, Yudong Wang and Xu Wang
Plants 2024, 13(9), 1276; https://doi.org/10.3390/plants13091276 - 05 May 2024
Abstract
Deep learning has emerged as a powerful tool for investigating intricate biological processes in plants by harnessing the potential of large-scale data. Gene regulation is a complex process that transcription factors (TFs), cooperating with their target genes, participate in through various aspects of
[...] Read more.
Deep learning has emerged as a powerful tool for investigating intricate biological processes in plants by harnessing the potential of large-scale data. Gene regulation is a complex process that transcription factors (TFs), cooperating with their target genes, participate in through various aspects of biological processes. Despite its significance, the study of gene regulation has primarily focused on a limited number of notable instances, leaving numerous aspects and interactions yet to be explored comprehensively. Here, we developed DEGRN (Deep learning on Expression for Gene Regulatory Network), an innovative deep learning model designed to decipher gene interactions by leveraging high-dimensional expression data obtained from bulk RNA-Seq and scRNA-Seq data in the model plant Arabidopsis. DEGRN exhibited a compared level of predictive power when applied to various datasets. Through the utilization of DEGRN, we successfully identified an extensive set of 3,053,363 high-quality interactions, encompassing 1430 TFs and 13,739 non-TF genes. Notably, DEGRN’s predictive capabilities allowed us to uncover novel regulators involved in a range of complex biological processes, including development, metabolism, and stress responses. Using leaf senescence as an example, we revealed a complex network underpinning this process composed of diverse TF families, including bHLH, ERF, and MYB. We also identified a novel TF, named MAF5, whose expression showed a strong linear regression relation during the progression of senescence. The mutant maf5 showed early leaf decay compared to the wild type, indicating a potential role in the regulation of leaf senescence. This hypothesis was further supported by the expression patterns observed across four stages of leaf development, as well as transcriptomics analysis. Overall, the comprehensive coverage provided by DEGRN expands our understanding of gene regulatory networks and paves the way for further investigations into their functional implications.
Full article
(This article belongs to the Section Plant Genetics, Genomics and Biotechnology)
►▼
Show Figures
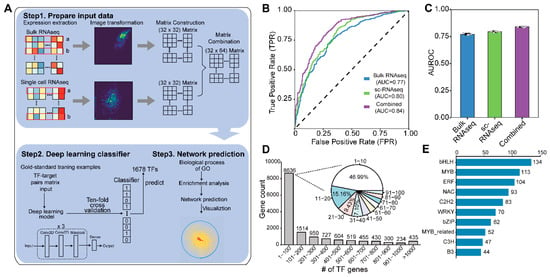
Figure 1
Open AccessArticle
Genetic Dissection of Diverse Seed Coat Patterns in Cowpea through a Comprehensive GWAS Approach
by
Haizheng Xiong, Yilin Chen, Waltram Ravelombola, Beiquan Mou, Xiaolun Sun, Qingyang Zhang, Yiting Xiao, Yang Tian, Qun Luo, Ibtisam Alatawi, Kenani Edward Chiwina, Hanan Mohammedsaeed Alkabkabi and Ainong Shi
Plants 2024, 13(9), 1275; https://doi.org/10.3390/plants13091275 - 05 May 2024
Abstract
This study investigates the genetic determinants of seed coat color and pattern variations in cowpea (Vigna unguiculata), employing a genome-wide association approach. Analyzing a mapping panel of 296 cowpea varieties with 110,000 single nucleotide polymorphisms (SNPs), we focused on eight unique
[...] Read more.
This study investigates the genetic determinants of seed coat color and pattern variations in cowpea (Vigna unguiculata), employing a genome-wide association approach. Analyzing a mapping panel of 296 cowpea varieties with 110,000 single nucleotide polymorphisms (SNPs), we focused on eight unique coat patterns: (1) Red and (2) Cream seed; (3) White and (4) Brown/Tan seed coat; (5) Pink, (6) Black, (7) Browneye and (8) Red/Brown Holstein. Across six GWAS models (GLM, SRM, MLM, MLMM, FarmCPU from GAPIT3, and TASSEL5), 13 significant SNP markers were identified and led to the discovery of 23 candidate genes. Among these, four specific genes may play a direct role in determining seed coat pigment. These findings lay a foundational basis for future breeding programs aimed at creating cowpea varieties aligned with consumer preferences and market requirements.
Full article
(This article belongs to the Special Issue Genetic Diversity of Germplasm Resources in Cereals and Legumes)
►▼
Show Figures

Figure 1
Open AccessArticle
Genome-Wide Identification and Evolutionary Analysis of Receptor-like Kinase Family Genes Provides Insights into Anthracnose Resistance of Dioscorea alata
by
Yuqian Jiang, Xin-Yu Lu, Ya-Li Qin, Yan-Mei Zhang and Zhu-Qing Shao
Plants 2024, 13(9), 1274; https://doi.org/10.3390/plants13091274 - 05 May 2024
Abstract
Dioscorea alata, commonly known as “greater yam”, is a vital crop in tropical and subtropical regions of the world, yet it faces significant threats from anthracnose disease, mainly caused by Colletotrichum gloeosporioides. However, exploring disease resistance genes in this species has
[...] Read more.
Dioscorea alata, commonly known as “greater yam”, is a vital crop in tropical and subtropical regions of the world, yet it faces significant threats from anthracnose disease, mainly caused by Colletotrichum gloeosporioides. However, exploring disease resistance genes in this species has been challenging due to the difficulty of genetic mapping resulting from the loss of the flowering trait in many varieties. The receptor-like kinase (RLK) gene family represents essential immune receptors in plants. In this study, genomic analysis revealed 467 RLK genes in D. alata. The identified RLKs were distributed unevenly across chromosomes, likely due to tandem duplication events. However, a considerable number of ancient whole-genome or segmental duplications dating back over 100 million years contributed to the diversity of RLK genes. Phylogenetic analysis unveiled at least 356 ancient RLK lineages in the common ancestor of Dioscoreaceae, which differentially inherited and expanded to form the current RLK profiles of D. alata and its relatives. The analysis of cis-regulatory elements indicated the involvement of RLK genes in diverse stress responses. Transcriptome analysis identified RLKs that were up-regulated in response to C. gloeosporioides infection, suggesting their potential role in resisting anthracnose disease. These findings provide novel insights into the evolution of RLK genes in D. alata and their potential contribution to disease resistance.
Full article
(This article belongs to the Special Issue Origin, Evolution and Functional Mechanisms of Plant Immune System)
►▼
Show Figures
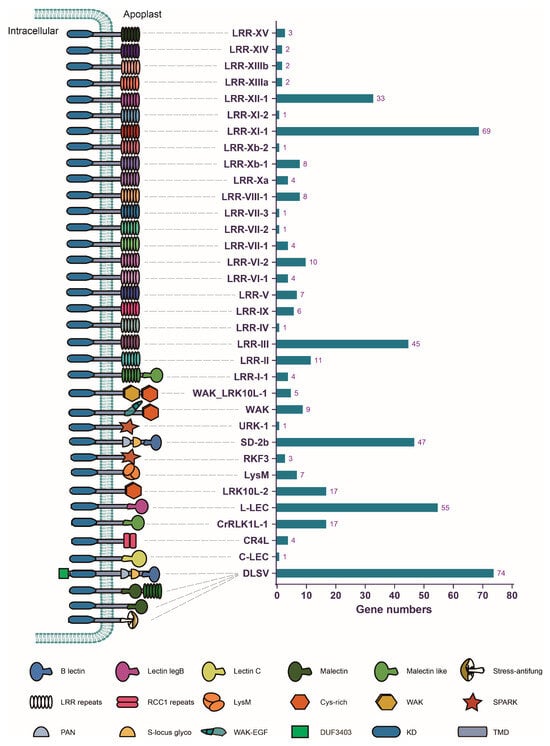
Figure 1
Open AccessArticle
Identification of Key Genes of Fruit Shape Variation in Jujube with Integrating Elliptic Fourier Descriptors and Transcriptome
by
Yue Ren, Wenqing Fu, Yi Gao, Yuhan Chen, Decang Kong, Ming Cao, Xiaoming Pang and Wenhao Bo
Plants 2024, 13(9), 1273; https://doi.org/10.3390/plants13091273 - 05 May 2024
Abstract
Jujube (Ziziphus jujuba) exhibits a rich diversity in fruit shape, with natural occurrences of gourd-like, flattened, and other special shapes. Despite the ongoing research into fruit shape, studies integrating elliptical Fourier descriptors (EFDs) with both Short Time-series Expression Miner (STEM) and
[...] Read more.
Jujube (Ziziphus jujuba) exhibits a rich diversity in fruit shape, with natural occurrences of gourd-like, flattened, and other special shapes. Despite the ongoing research into fruit shape, studies integrating elliptical Fourier descriptors (EFDs) with both Short Time-series Expression Miner (STEM) and weighted gene co-expression network analysis (WGCNA) for gene discovery remain scarce. In this study, six cultivars of jujube fruits with distinct shapes were selected, and samples were collected from the fruit set period to the white mature stage across five time points for shape analysis and transcriptome studies. By combining EFDs with WGCNA and STEM, the study aimed to identify the critical periods and key genes involved in the formation of jujube fruit shape. The findings indicated that the D25 (25 days after flowering) is crucial for the development of jujube fruit shape. Moreover, ZjAGL80, ZjABI3, and eight other genes have been implicated to regulate the shape development of jujubes at different periods of fruit development, through seed development and fruit development pathway. In this research, EFDs were employed to precisely delineate the shape of jujube fruits. This approach, in conjunction with transcriptome, enhanced the precision of gene identification, and offered an innovative methodology for fruit shape analysis. This integration facilitates the advancement of research into the morphological characteristics of plant fruits, underpinning the development of a refined framework for the genetic underpinnings of fruit shape variation.
Full article
(This article belongs to the Special Issue Genetic Breeding of Trees)
►▼
Show Figures

Figure 1
Open AccessReview
Application of Developmental Regulators for Enhancing Plant Regeneration and Genetic Transformation
by
Pingjun Xu, Yinxiao Zhong, Ang Xu, Bingshuang Liu, Yue Zhang, Anqi Zhao, Xiaoming Yang, Meiling Ming, Fuliang Cao and Fangfang Fu
Plants 2024, 13(9), 1272; https://doi.org/10.3390/plants13091272 - 04 May 2024
Abstract
►▼
Show Figures
Establishing plant regeneration systems and efficient genetic transformation techniques plays a crucial role in plant functional genomics research and the development of new crop varieties. The inefficient methods of transformation and regeneration of recalcitrant species and the genetic dependence of the transformation process
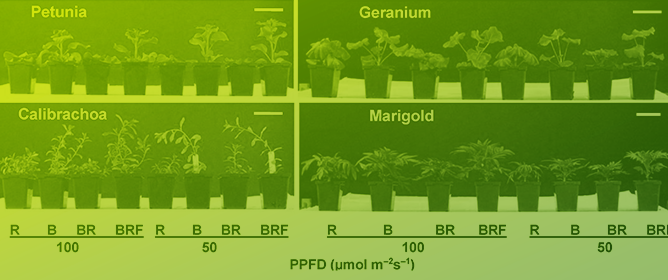
[...] Read more.
Establishing plant regeneration systems and efficient genetic transformation techniques plays a crucial role in plant functional genomics research and the development of new crop varieties. The inefficient methods of transformation and regeneration of recalcitrant species and the genetic dependence of the transformation process remain major obstacles. With the advancement of plant meristematic tissues and somatic embryogenesis research, several key regulatory genes, collectively known as developmental regulators, have been identified. In the field of plant genetic transformation, the application of developmental regulators has recently garnered significant interest. These regulators play important roles in plant growth and development, and when applied in plant genetic transformation, they can effectively enhance the induction and regeneration capabilities of plant meristematic tissues, thus providing important opportunities for improving genetic transformation efficiency. This review focuses on the introduction of several commonly used developmental regulators. By gaining an in-depth understanding of and applying these developmental regulators, it is possible to further enhance the efficiency and success rate of plant genetic transformation, providing strong support for plant breeding and genetic engineering research.
Full article
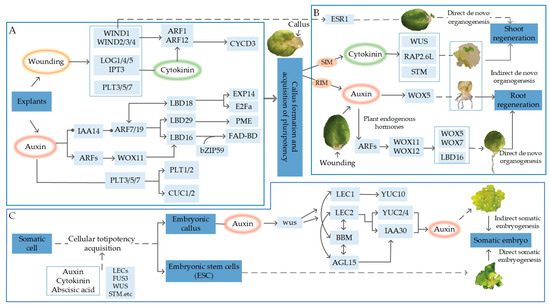
Figure 1
Open AccessArticle
Environmental and Biogeographic Drivers behind Alpine Plant Thermal Tolerance and Genetic Variation
by
Lisa M. Danzey, Verónica F. Briceño, Alicia M. Cook, Adrienne B. Nicotra, Gwendolyn Peyre, Maurizio Rossetto, Jia-Yee S. Yap and Andrea Leigh
Plants 2024, 13(9), 1271; https://doi.org/10.3390/plants13091271 - 04 May 2024
Abstract
In alpine ecosystems, elevation broadly functions as a steep thermal gradient, with plant communities exposed to regular fluctuations in hot and cold temperatures. These conditions lead to selective filtering, potentially contributing to species-level variation in thermal tolerance and population-level genetic divergence. Few studies
[...] Read more.
In alpine ecosystems, elevation broadly functions as a steep thermal gradient, with plant communities exposed to regular fluctuations in hot and cold temperatures. These conditions lead to selective filtering, potentially contributing to species-level variation in thermal tolerance and population-level genetic divergence. Few studies have explored the breadth of alpine plant thermal tolerances across a thermal gradient or the underlying genetic variation thereof. We measured photosystem heat (Tcrit-hot) and cold (Tcrit-cold) thresholds of ten Australian alpine species across elevation gradients and characterised their neutral genetic variation. To reveal the biogeographical drivers of present-day genetic signatures, we also reconstructed temporal changes in habitat suitability across potential distributional ranges. We found intraspecific variation in thermal thresholds, but this was not associated with elevation, nor underpinned by genetic differentiation on a local scale. Instead, regional population differentiation and considerable homozygosity within populations may, in part, be driven by distributional contractions, long-term persistence, and migrations following habitat suitability. Our habitat suitability models suggest that cool-climate-distributed alpine plants may be threatened by a warming climate. Yet, the observed wide thermal tolerances did not reflect this vulnerability. Conservation efforts should seek to understand variations in species-level thermal tolerance across alpine microclimates.
Full article
(This article belongs to the Section Plant Ecology)
►▼
Show Figures

Figure 1

Journal Menu
► ▼ Journal Menu-
- Plants Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Agronomy, Beverages, Fermentation, Horticulturae, Plants
Grapevine Facing Climate Change: From Land, through Plants to Grapes and Wine
Topic Editors: Othmane Merah, Ana Fernandes De Oliveira, Daniela Satta, Mario Cunha, Jesus Yuste, Jalloul BouajilaDeadline: 30 June 2024
Topic in
Agronomy, Diversity, Forests, IJPB, Plants
Plant Invasion
Topic Editors: Bruce Osborne, Panayiotis G. DimitrakopoulosDeadline: 31 July 2024
Topic in
Agriculture, Agronomy, Microorganisms, Plants, Soil Systems, Nitrogen
Carbon and Nitrogen Cycling in Agro-Ecosystems and Other Anthropogenically Maintained Ecosystems
Topic Editors: Jie Li, Adnan Mustafa, Jan FrouzDeadline: 30 September 2024
Topic in
Agronomy, Crops, Plants
Abiotic Stress Responses in Wheat: Perspectives on Productivity and Sustainability
Topic Editors: Wenshan Guo, Jinfeng Ding, Min ZhuDeadline: 31 October 2024

Conferences
Special Issues
Special Issue in
Plants
Structural and Functional Analysis of Extracts in Plants IV
Guest Editor: Stefania LamponiDeadline: 10 May 2024
Special Issue in
Plants
Traditional Cultivars as a Genetic Source of Stress Tolerance and Quality Enhancement
Guest Editors: Marija Viljevac Vuletić, Ines MihaljevićDeadline: 25 May 2024
Special Issue in
Plants
Diagnosis and Control of Plant Bacterial Diseases
Guest Editor: Joel L. VannesteDeadline: 31 May 2024
Special Issue in
Plants
Microscopy Techniques in Plant Studies
Guest Editors: Emilio de Castro Miguel, Maura Da Cunha, Thaiz Batista Azevedo Rangel MiguelDeadline: 20 June 2024
Topical Collections
Topical Collection in
Plants
Advances in Plant Diversification and Biosystematics
Collection Editors: Yang Liu, Ângela Sartori
Topical Collection in
Plants
New Trends in Plant Science in Italy
Collection Editors: Claudio Moser, Massimo Galbiati
Topical Collection in
Plants
New Trends in Plant Science in China
Collection Editors: Ming Chen, Dijun Chen, Xianwen Meng
Topical Collection in
Plants
Essential Oils of Plants (Chemical Composition, Variation and Properties)
Collection Editors: Jésus Palá-Pául, Joe Brophy











