-
 Neuroprotective Effects of the Nutraceutical Dehydrozingerone and Its C2-Symmetric Dimer in a Drosophila Model of Parkinson’s Disease
Neuroprotective Effects of the Nutraceutical Dehydrozingerone and Its C2-Symmetric Dimer in a Drosophila Model of Parkinson’s Disease -
 Role of DNase Activity in Human Sperm DNA Fragmentation
Role of DNase Activity in Human Sperm DNA Fragmentation -
 Volaboloma Mortis Fingerprint for Post-Mortem Interval Estimation
Volaboloma Mortis Fingerprint for Post-Mortem Interval Estimation -
 Exploring FDA-Approved Frontiers: Insights into Natural and Engineered Peptide Analogues
Exploring FDA-Approved Frontiers: Insights into Natural and Engineered Peptide Analogues
Journal Description
Biomolecules
Biomolecules
is a peer-reviewed, open access journal on structures and functions of bioactive and biogenic substances, molecular mechanisms with biological and medical implications as well as biomaterials and their applications. Biomolecules is published monthly online by MDPI. The Spanish Society for Biochemistry and Molecular Biology (SEBBM) is affiliated with Biomolecules and their members receive discounts on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, MEDLINE, PMC, Embase, CAPlus / SciFinder, and other databases.
- Journal Rank: JCR - Q1 (Biochemistry & Molecular Biology) / CiteScore - Q1 (Biochemistry)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.9 days after submission; acceptance to publication is undertaken in 2.9 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Sections: published in 19 topical sections.
- Testimonials: See what our editors and authors say about Biomolecules.
- Companion journal: Receptors.
Impact Factor:
5.5 (2022);
5-Year Impact Factor:
5.8 (2022)
Latest Articles
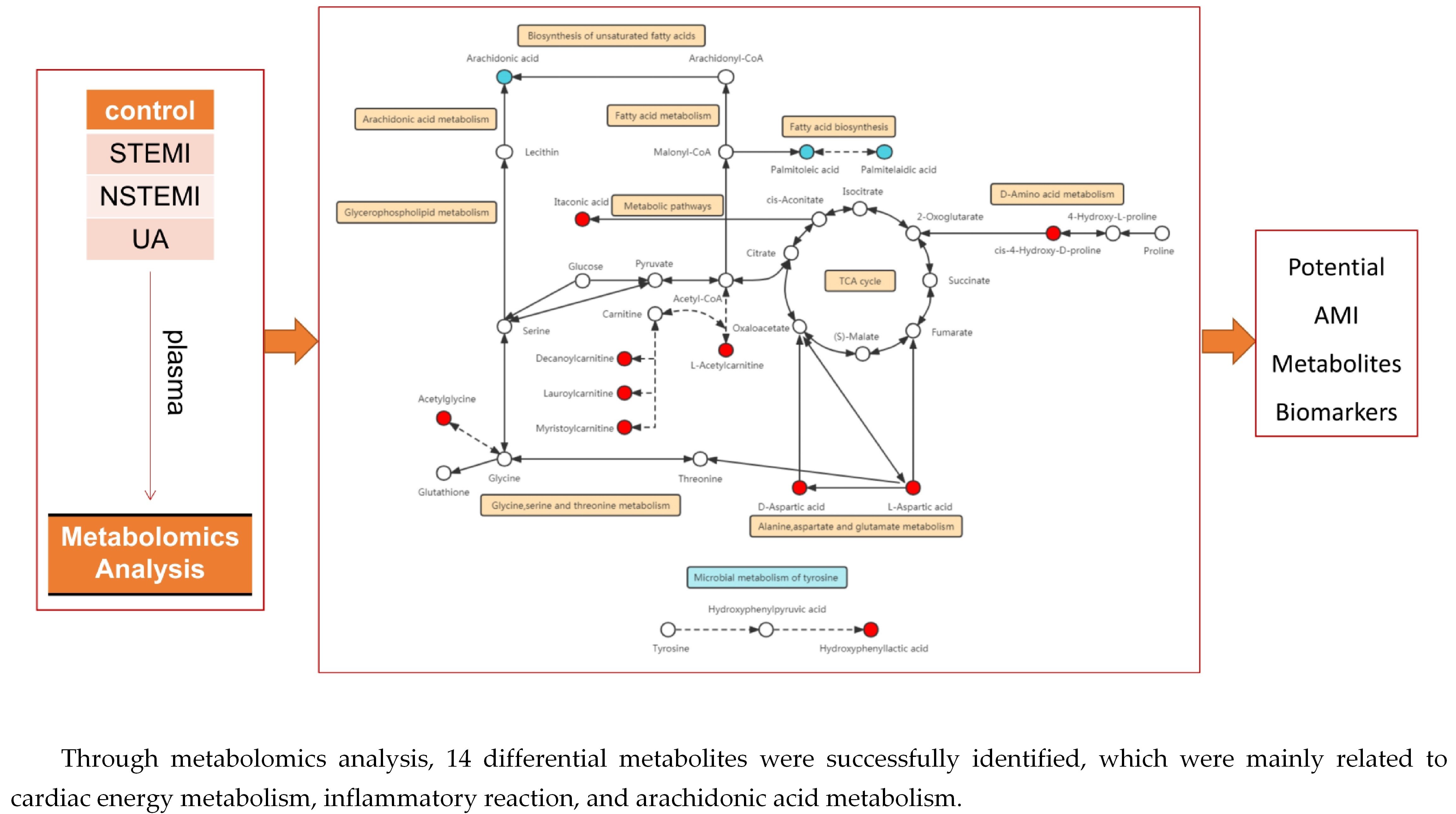
Metabolomics Analysis Identifies Differential Metabolites as Biomarkers for Acute Myocardial Infarction
Biomolecules 2024, 14(5), 532; https://doi.org/10.3390/biom14050532 (registering DOI) - 29 Apr 2024
Abstract
Myocardial infarction (MI), including ST-segment elevation MI (STEMI) and non-ST-segment elevation MI (NSTEMI), is still a leading cause of death worldwide. Metabolomics technology was used to explore differential metabolites (DMs) as potential biomarkers for early diagnosis of STEMI and NSTEMI. In the study,
[...] Read more.
Myocardial infarction (MI), including ST-segment elevation MI (STEMI) and non-ST-segment elevation MI (NSTEMI), is still a leading cause of death worldwide. Metabolomics technology was used to explore differential metabolites (DMs) as potential biomarkers for early diagnosis of STEMI and NSTEMI. In the study, 2531 metabolites, including 1925 DMs, were discovered. In the selected 27 DMs, 14 were successfully verified in a new cohort, and the AUC values were all above 0.8. There were 10 in STEMI group, namely L-aspartic acid, L-acetylcarnitine, acetylglycine, decanoylcarnitine, hydroxyphenyllactic acid, ferulic acid, itaconic acid, lauroylcarnitine, myristoylcarnitine, and cis-4-hydroxy-D-proline, and 5 in NSTEMI group, namely L-aspartic acid, arachidonic acid, palmitoleic acid, D-aspartic acid, and palmitelaidic acid. These 14 DMs may be developed as biomarkers for the early diagnosis of MI with high sensitivity and specificity. These findings have particularly important clinical significance for NSTEMI patients because these patients have no typical ECG changes.
Full article
(This article belongs to the Special Issue Molecular Biomarkers In Cardiology 2022–2023)
►
Show Figures
Open AccessArticle
Kinetics of Human Serum Albumin Adsorption on Polycation Functionalized Silica
by
Małgorzata Nattich-Rak, Dominik Kosior, Maria Morga and Zbigniew Adamczyk
Biomolecules 2024, 14(5), 531; https://doi.org/10.3390/biom14050531 (registering DOI) - 29 Apr 2024
Abstract
The adsorption kinetics of human serum albumin (HSA) on bare and poly-L-arginine (PARG)-modified silica substrates were investigated using reflectometry and atomic force microscopy (AFM). Measurements were carried out at various pHs, flow rates and albumin concentrations in the 10 and 150 mM NaCl
[...] Read more.
The adsorption kinetics of human serum albumin (HSA) on bare and poly-L-arginine (PARG)-modified silica substrates were investigated using reflectometry and atomic force microscopy (AFM). Measurements were carried out at various pHs, flow rates and albumin concentrations in the 10 and 150 mM NaCl solutions. The mass transfer rate constants and the maximum protein coverages were determined for the bare silica at pH 4.0 and theoretically interpreted in terms of the hybrid random sequential adsorption model. These results were used as reference data for the analysis of adsorption kinetics at larger pHs. It was shown that the adsorption on bare silica rapidly decreased with pH and became negligible at pH 7.4. The albumin adsorption on PARG-functionalized silica showed an opposite trend, i.e., it was negligible at pH 4 and attained maximum values at pH 7.4 and 150 mM NaCl, the conditions corresponding to the blood serum environment. These results were interpreted as the evidence of a significant role of electrostatic interactions in the albumin adsorption on the bare and PARG-modified silica. It was also argued that our results can serve as useful reference data enabling a proper interpretation of protein adsorption on substrates functionalized by polyelectrolytes.
Full article
(This article belongs to the Special Issue Mechanisms of Formation of Biomolecule Complexes: Theory, Experiment, Applications)
Open AccessReview
Diruthenium Paddlewheel Complexes Attacking Proteins: Axial Versus Equatorial Coordination
by
Iogann Tolbatov, Paolo Umari and Alessandro Marrone
Biomolecules 2024, 14(5), 530; https://doi.org/10.3390/biom14050530 (registering DOI) - 28 Apr 2024
Abstract
Metallodrugs are an important group of medicinal agents used for the treatment of various diseases ranging from cancers to viral, bacterial, and parasitic diseases. Their distinctive features include the availability of a metal centre, redox activity, as well as the ability to multitarget.
[...] Read more.
Metallodrugs are an important group of medicinal agents used for the treatment of various diseases ranging from cancers to viral, bacterial, and parasitic diseases. Their distinctive features include the availability of a metal centre, redox activity, as well as the ability to multitarget. Diruthenium paddlewheel complexes are an intensely developing group of metal scaffolds, which can securely coordinate bidentate xenobiotics and transport them to target tissues, releasing them by means of substitution reactions with biomolecular nucleophiles. It is of the utmost importance to gain a complete comprehension of which chemical reactions happen with them in physiological milieu to design novel drugs based on these bimetallic scaffolds. This review presents the data obtained in experiments and calculations, which clarify the chemistry these complexes undergo once administered in the proteic environment. This study demonstrates how diruthenium paddlewheel complexes may indeed embody a new paradigm in the design of metal-based drugs of dual-action by presenting and discussing the protein metalation by these complexes.
Full article
Open AccessReview
Unlocking Genetic Mysteries during the Epic Sperm Journey toward Fertilization: Further Expanding Cre Mouse Lines
by
Pengyuan Dai, Chaoye Ma, Chen Chen, Min Liang, Shijue Dong, Hao Chen and Xiaoning Zhang
Biomolecules 2024, 14(5), 529; https://doi.org/10.3390/biom14050529 (registering DOI) - 28 Apr 2024
Abstract
►▼
Show Figures
The spatiotemporal expression patterns of genes are crucial for maintaining normal physiological functions in animals. Conditional gene knockout using the cyclization recombination enzyme (Cre)/locus of crossover of P1 (Cre/LoxP) strategy has been extensively employed for functional assays
[...] Read more.
The spatiotemporal expression patterns of genes are crucial for maintaining normal physiological functions in animals. Conditional gene knockout using the cyclization recombination enzyme (Cre)/locus of crossover of P1 (Cre/LoxP) strategy has been extensively employed for functional assays at specific tissue or developmental stages. This approach aids in uncovering the associations between phenotypes and gene regulation while minimizing interference among distinct tissues. Various Cre-engineered mouse models have been utilized in the male reproductive system, including Dppa3-MERCre for primordial germ cells, Ddx4-Cre and Stra8-Cre for spermatogonia, Prm1-Cre and Acrv1-iCre for haploid spermatids, Cyp17a1-iCre for the Leydig cell, Sox9-Cre for the Sertoli cell, and Lcn5/8/9-Cre for differentiated segments of the epididymis. Notably, the specificity and functioning stage of Cre recombinases vary, and the efficiency of recombination driven by Cre depends on endogenous promoters with different sequences as well as the constructed Cre vectors, even when controlled by an identical promoter. Cre mouse models generated via traditional recombination or CRISPR/Cas9 also exhibit distinct knockout properties. This review focuses on Cre-engineered mouse models applied to the male reproductive system, including Cre-targeting strategies, mouse model screening, and practical challenges encountered, particularly with novel mouse strains over the past decade. It aims to provide valuable references for studies conducted on the male reproductive system.
Full article

Figure 1
Open AccessReview
NMR Spectroscopy in Diagnosis and Monitoring of Methylmalonic and Propionic Acidemias
by
Calin Deleanu and Alina Nicolescu
Biomolecules 2024, 14(5), 528; https://doi.org/10.3390/biom14050528 (registering DOI) - 28 Apr 2024
Abstract
Although both localized nuclear magnetic resonance spectroscopy (MRS) and non-localized nuclear magnetic resonance spectroscopy (NMR) generate the same information, i.e., spectra generated by various groups from the structure of metabolites, they are rarely employed in the same study or by the same research
[...] Read more.
Although both localized nuclear magnetic resonance spectroscopy (MRS) and non-localized nuclear magnetic resonance spectroscopy (NMR) generate the same information, i.e., spectra generated by various groups from the structure of metabolites, they are rarely employed in the same study or by the same research group. As our review reveals, these techniques have never been applied in the same study of methylmalonic acidemia (MMA), propionic acidemia (PA) or vitamin B12 deficiency patients. On the other hand, MRS and NMR provide complementary information which is very valuable in the assessment of the severity of disease and efficiency of its treatment. Thus, MRS provides intracellular metabolic information from localized regions of the brain, while NMR provides extracellular metabolic information from biological fluids like urine, blood or cerebrospinal fluid. This paper presents an up-to-date review of the NMR and MRS studies reported to date for methylmalonic and propionic acidemias. Vitamin B12 deficiency, although in most of its cases not inherited, shares similarities in its metabolic effects with MMA and it is also covered in this review.
Full article
(This article belongs to the Collection Metabolomics and Integrated Multi-Omics in Health and Disease)
►▼
Show Figures
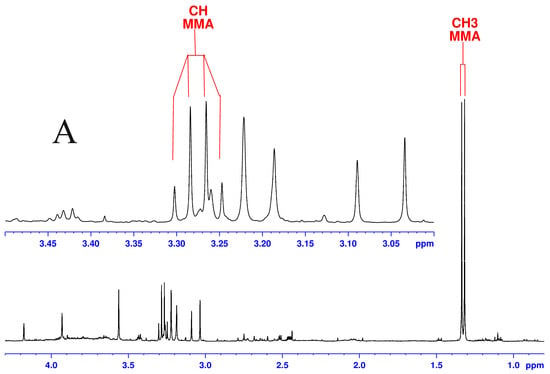
Figure 1
Open AccessArticle
The Influence of Stress and Binge-Patterned Alcohol Drinking on Mouse Skeletal Muscle Protein Synthesis and Degradation Pathways
by
Carter H Reed, Anna C. Tystahl, Hyeyoon Eo, Trevor J. Buhr, Ella E. Bauer, Ji Heun Lee, Peter J. Clark and Rudy J. Valentine
Biomolecules 2024, 14(5), 527; https://doi.org/10.3390/biom14050527 (registering DOI) - 28 Apr 2024
Abstract
Adverse experiences (e.g., acute stress) and alcohol misuse can both impair skeletal muscle homeostasis, resulting in reduced protein synthesis and greater protein breakdown. Exposure to acute stress is a significant risk factor for engaging in alcohol misuse. However, little is known about how
[...] Read more.
Adverse experiences (e.g., acute stress) and alcohol misuse can both impair skeletal muscle homeostasis, resulting in reduced protein synthesis and greater protein breakdown. Exposure to acute stress is a significant risk factor for engaging in alcohol misuse. However, little is known about how these factors together might further affect skeletal muscle health. To that end, this study investigated the effects of acute stress exposure followed by a period of binge-patterned alcohol drinking on signaling factors along mouse skeletal muscle protein synthesis (MPS) and degradation (MPD) pathways. Young adult male C57BL/6J mice participated in the Drinking in the Dark paradigm, where they received 2–4 h of access to 20% ethanol (alcohol group) or water (control group) for four days to establish baseline drinking levels. Three days later, half of the mice in each group were either exposed to a single episode of uncontrollable tail shocks (acute stress) or remained undisturbed in their home cages (no stress). Three days after stress exposure, mice received 4 h of access to 20% ethanol (alcohol) to model binge-patterned alcohol drinking or water for ten consecutive days. Immediately following the final episode of alcohol access, mouse gastrocnemius muscle was extracted to measure changes in relative protein levels along the Akt-mTOR MPS, as well as the ubiquitin-proteasome pathway (UPP) and autophagy MPD pathways via Western blotting. A single exposure to acute stress impaired Akt singling and reduced rates of MPS, independent of alcohol access. This observation was concurrent with a potent increase in heat shock protein seventy expression in the muscle of stressed mice. Alcohol drinking did not exacerbate stress-induced alterations in the MPS and MPD signaling pathways. Instead, changes in the MPS and MPD signaling factors due to alcohol access were primarily observed in non-stressed mice. Taken together, these data suggest that exposure to a stressor of sufficient intensity may cause prolonged disruptions to signaling factors that impact skeletal muscle health and function beyond what could be further induced by periods of alcohol misuse.
Full article
(This article belongs to the Special Issue Skeletal Muscle Homeostasis and Regeneration)
►▼
Show Figures
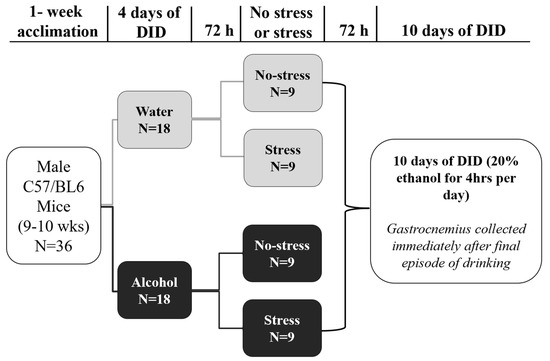
Figure 1
Open AccessFeature PaperArticle
Synthesis of the Antimicrobial Peptide Murepavadin Using Novel Coupling Agents
by
Júlia García-Gros, Yolanda Cajal, Ana Maria Marqués and Francesc Rabanal
Biomolecules 2024, 14(5), 526; https://doi.org/10.3390/biom14050526 (registering DOI) - 27 Apr 2024
Abstract
The problem of antimicrobial resistance is becoming a daunting challenge for human society and healthcare systems around the world. Hence, there is a constant need to develop new antibiotics to fight resistant bacteria, among other important social and economic measures. In this regard,
[...] Read more.
The problem of antimicrobial resistance is becoming a daunting challenge for human society and healthcare systems around the world. Hence, there is a constant need to develop new antibiotics to fight resistant bacteria, among other important social and economic measures. In this regard, murepavadin is a cyclic antibacterial peptide in development. The synthesis of murepavadin was undertaken in order to optimize the preparative protocol and scale-up, in particular, the use of new activation reagents. In our hands, classical approaches using carbodiimide/hydroxybenzotriazole rendered low yields. The use of novel carbodiimide and reagents based on OxymaPure® and Oxy-B is discussed together with the proper use of chromatographic conditions for the adequate characterization of peptide crudes. Higher yields and purities were obtained. Finally, the antimicrobial activity of different synthetic batches was tested in three Pseudomonas aeruginosa strains, including highly resistant ones. All murepavadin batches yielded the same highly active MIC values and proved that the chiral integrity of the molecule was preserved throughout the whole synthetic procedure.
Full article
(This article belongs to the Special Issue Development, Characterization, and Application of Bioactive Peptides, Diagnostic Biomarkers, and Pharmaceutical Proteins)
►▼
Show Figures
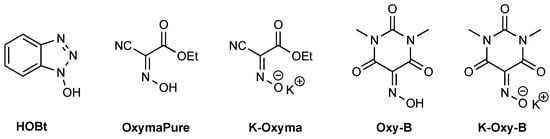
Figure 1
Open AccessArticle
Alpha-Melanocyte-Stimulating Hormone Maintains Retinal Homeostasis after Ischemia/Reperfusion
by
Tat Fong Ng, Jenna Y. Cho, John L. Zhao, John R. Gardiner, Eric S. Wang, Elman Leung, Ziqian Xu, Samantha L. Fineman, Melinda Lituchy, Amy C. Lo and Andrew W. Taylor
Biomolecules 2024, 14(5), 525; https://doi.org/10.3390/biom14050525 (registering DOI) - 27 Apr 2024
Abstract
Augmenting the natural melanocortin pathway in mouse eyes with uveitis or diabetes protects the retinas from degeneration. The retinal cells are protected from oxidative and apoptotic signals of death. Therefore, we investigated the effects of a therapeutic application of the melanocortin alpha-melanocyte-stimulating hormone
[...] Read more.
Augmenting the natural melanocortin pathway in mouse eyes with uveitis or diabetes protects the retinas from degeneration. The retinal cells are protected from oxidative and apoptotic signals of death. Therefore, we investigated the effects of a therapeutic application of the melanocortin alpha-melanocyte-stimulating hormone (α-MSH) on an ischemia and reperfusion (I/R) model of retinal degenerative disease. Eyes were subjected to an I/R procedure and were treated with α-MSH. Retinal sections were histopathologically scored. Also, the retinal sections were immunostained for viable ganglion cells, activated Muller cells, microglial cells, and apoptosis. The I/R caused retinal deformation and ganglion cell loss that was significantly reduced in I/R eyes treated with α-MSH. While α-MSH treatment marginally reduced the number of GFAP-positive Muller cells, it significantly suppressed the density of Iba1-positive microglial cells in the I/R retinas. Within one hour after I/R, there was apoptosis in the ganglion cell layer, and by 48 h, there was apoptosis in all layers of the neuroretina. The α-MSH treatment significantly reduced and delayed the onset of apoptosis in the retinas of I/R eyes. The results demonstrate that therapeutically augmenting the melanocortin pathways preserves retinal structure and cell survival in eyes with progressive neuroretinal degenerative disease.
Full article
(This article belongs to the Special Issue Diabetic Retinopathy: From Pathophysiology to Therapeutic Perspectives)
►▼
Show Figures
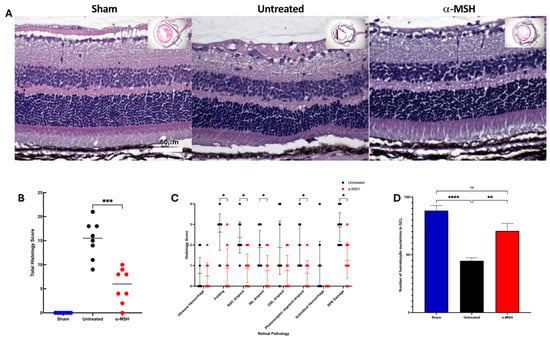
Figure 1
Open AccessArticle
Unraveling the Etiology of Dilated Cardiomyopathy through Differential miRNA–mRNA Interactome
by
Fernando Bonet, Francisco Hernandez-Torres, Mónica Ramos-Sánchez, Maribel Quezada-Feijoo, Aníbal Bermúdez-García, Tomás Daroca, Elena Alonso-Villa, Carlos García-Padilla, Alipio Mangas and Rocio Toro
Biomolecules 2024, 14(5), 524; https://doi.org/10.3390/biom14050524 (registering DOI) - 27 Apr 2024
Abstract
Dilated cardiomyopathy (DCM) encompasses various acquired or genetic diseases sharing a common phenotype. The understanding of pathogenetic mechanisms and the determination of the functional effects of each etiology may allow for tailoring different therapeutic strategies. MicroRNAs (miRNAs) have emerged as key regulators in
[...] Read more.
Dilated cardiomyopathy (DCM) encompasses various acquired or genetic diseases sharing a common phenotype. The understanding of pathogenetic mechanisms and the determination of the functional effects of each etiology may allow for tailoring different therapeutic strategies. MicroRNAs (miRNAs) have emerged as key regulators in cardiovascular diseases, including DCM. However, their specific roles in different DCM etiologies remain elusive. Here, we applied mRNA-seq and miRNA-seq to identify the gene and miRNA signature from myocardial biopsies from four patients with DCM caused by volume overload (VCM) and four with ischemic DCM (ICM). Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis were used for differentially expressed genes (DEGs). The miRNA–mRNA interactions were identified by Pearson correlation analysis and miRNA target-prediction programs. mRNA-seq and miRNA-seq were validated by qRT-PCR and miRNA–mRNA interactions were validated by luciferase assays. We found 112 mRNAs and five miRNAs dysregulated in VCM vs. ICM. DEGs were positively enriched for pathways related to the extracellular matrix (ECM), mitochondrial respiration, cardiac muscle contraction, and fatty acid metabolism in VCM vs. ICM and negatively enriched for immune-response-related pathways, JAK-STAT, and NF-kappa B signaling. We identified four pairs of negatively correlated miRNA–mRNA: miR-218-5p-DDX6, miR-218-5p-TTC39C, miR-218-5p-SEMA4A, and miR-494-3p-SGMS2. Our study revealed novel miRNA–mRNA interaction networks and signaling pathways for VCM and ICM, providing novel insights into the development of these DCM etiologies.
Full article
(This article belongs to the Section Molecular Genetics)
Open AccessArticle
Herpesvirus Entry Mediator as an Immune Checkpoint Target and a Potential Prognostic Biomarker in Myeloid and Lymphoid Leukemia
by
Fatemah S. Basingab, Reem A. Alzahrani, Aisha A. Alrofaidi, Ahmed S. Barefah, Rawan M. Hammad, Hadil M. Alahdal, Jehan S. Alrahimi, Kawther A. Zaher, Ali H. Algiraigri, Mai M. El-Daly, Saleh A. Alkarim and Alia M. Aldahlawi
Biomolecules 2024, 14(5), 523; https://doi.org/10.3390/biom14050523 (registering DOI) - 27 Apr 2024
Abstract
Herpesvirus entry mediator (HVEM) is a molecular switch that can modulate immune responses against cancer. The significance of HVEM as an immune checkpoint target and a potential prognostic biomarker in malignancies is still controversial. This study aims to determine whether HVEM is an
[...] Read more.
Herpesvirus entry mediator (HVEM) is a molecular switch that can modulate immune responses against cancer. The significance of HVEM as an immune checkpoint target and a potential prognostic biomarker in malignancies is still controversial. This study aims to determine whether HVEM is an immune checkpoint target with inhibitory effects on anti-tumor CD4+ T cell responses in vitro and whether HVEM gene expression is dysregulated in patients with acute lymphocytic leukemia (ALL). HVEM gene expression in tumor cell lines and peripheral blood mononuclear cells (PBMCs) from ALL patients and healthy controls was measured using reverse transcription-quantitative polymerase chain reaction (RT-qPCR). Tumor cells were left untreated (control) or were treated with an HVEM blocker before co-culturing with CD4+ T cells in vitro in a carboxyfluorescein succinimidyl ester (CFSE)-dependent proliferation assay. HVEM expression was upregulated in the chronic myelogenous leukemia cell line (K562) (FC = 376.3, p = 0.086) compared with normal embryonic kidney cells (Hek293). CD4+ T cell proliferation was significantly increased in the HVEM blocker-treated K562 cells (p = 0.0033). Significant HVEM differences were detected in ALL PBMCs compared with the controls, and these were associated with newly diagnosed ALL (p = 0.0011) and relapsed/refractory (p = 0.0051) B cell ALL (p = 0.0039) patients. A significant differentiation between malignant ALL and the controls was observed in a receiver operating characteristic (ROC) curve analysis with AUC = 0.78 ± 0.092 (p = 0.014). These results indicate that HVEM is an inhibitory molecule that may serve as a target for immunotherapy and a potential ALL biomarker.
Full article
(This article belongs to the Topic MicroRNA: Mechanisms of Action, Physio-Pathological Implications, and Disease Biomarkers, 2nd Volume)
►▼
Show Figures
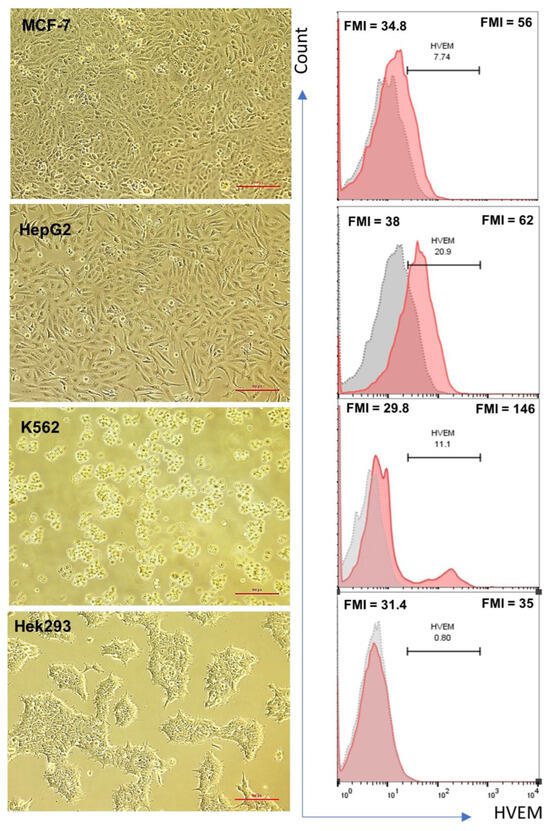
Figure 1
Open AccessArticle
The First Defined Null Allele of the Notch Regulator, a Suppressor of Deltex: Uncovering Its Novel Roles in Drosophila melanogaster Oogenesis
by
Marian B. Wilkin, Rory Whiteford, Tanveer Akbar, Samira Hosseini-Alghaderi, Raluca Revici, Ann-Marie Carbery and Martin Baron
Biomolecules 2024, 14(5), 522; https://doi.org/10.3390/biom14050522 (registering DOI) - 26 Apr 2024
Abstract
Suppressor of deltex (Su(dx)) is a Drosophila melanogaster member of the NEDD4 family of the HECT domain E3 ubiquitin ligases. Su(dx) acts as a regulator of Notch endocytic trafficking, promoting Notch lysosomal degradation and the down-regulation of both ligand-dependent and ligand-independent signalling, the
[...] Read more.
Suppressor of deltex (Su(dx)) is a Drosophila melanogaster member of the NEDD4 family of the HECT domain E3 ubiquitin ligases. Su(dx) acts as a regulator of Notch endocytic trafficking, promoting Notch lysosomal degradation and the down-regulation of both ligand-dependent and ligand-independent signalling, the latter involving trafficking through the endocytic pathway and activation of the endo/lysosomal membrane. Mutations of Su(dx) result in developmental phenotypes in the Drosophila wing that reflect increased Notch signalling, leading to gaps in the specification of the wing veins, and Su(dx) functions to provide the developmental robustness of Notch activity to environmental temperature shifts. The full developmental functions of Su(dx) are unclear; however, this is due to a lack of a clearly defined null allele. Here we report the first defined null mutation of Su(dx), generated by P-element excision, which removes the complete open reading frame. We show that the mutation is recessive-viable, with the Notch gain of function phenotypes affecting wing vein and leg development. We further uncover new roles for Su(dx) in Drosophila oogenesis, where it regulates interfollicular stalk formation, egg chamber separation and germline cyst enwrapment by the follicle stem cells. Interestingly, while the null allele exhibited a gain in Notch activity during oogenesis, the previously described Su(dx)SP allele, which carries a seven amino acid in-frame deletion, displayed a Notch loss of function phenotypes and an increase in follicle stem cell turnover. This is despite both alleles displaying similar Notch gain of function in wing development. We attribute this unexpected context-dependent outcome of Su(dx)sp being due to the partial retention of function by the intact C2 and WW domain regions of the protein. Our results extend our understanding of the developmental role of Su(dx) in the tissue renewal and homeostasis of the Drosophila ovary and illustrate the importance of examining an allelic series of mutations to fully understand developmental functions.
Full article
(This article belongs to the Special Issue Regulation of Notch Signaling Pathway and Its Relation to Diseases)
Open AccessArticle
Feature Matching of Microsecond-Pulsed Magnetic Fields Combined with Fe3O4 Particles for Killing A375 Melanoma Cells
by
Yan Mi, Meng-Nan Zhang, Chi Ma, Wei Zheng and Fei Teng
Biomolecules 2024, 14(5), 521; https://doi.org/10.3390/biom14050521 - 26 Apr 2024
Abstract
The combination of magnetic fields and magnetic nanoparticles (MNPs) to kill cancer cells by magneto-mechanical force represents a novel therapy, offering advantages such as non-invasiveness, among others. Pulsed magnetic fields (PMFs) hold promise for application in this therapy due to advantages such as
[...] Read more.
The combination of magnetic fields and magnetic nanoparticles (MNPs) to kill cancer cells by magneto-mechanical force represents a novel therapy, offering advantages such as non-invasiveness, among others. Pulsed magnetic fields (PMFs) hold promise for application in this therapy due to advantages such as easily adjustable parameters; however, they suffer from the drawback of narrow pulse width. In order to fully exploit the potential of PMFs and MNPs in this therapy, while maximizing therapeutic efficacy within the constraints of the narrow pulse width, a feature-matching theory is proposed, encompassing the matching of three aspects: (1) MNP volume and critical volume of Brownian relaxation, (2) relaxation time and pulse width, and (3) MNP shape and the intermittence of PMF. In the theory, a microsecond-PMF generator was developed, and four kinds of MNPs were selected for in vitro cell experiments. The results demonstrate that the killing rate of the experimental group meeting the requirements of the theory is at least 18% higher than the control group. This validates the accuracy of our theory and provides valuable guidance for the further application of PMFs in this therapy.
Full article
(This article belongs to the Special Issue Advanced Nanotechnology for Health and Diseases)
Open AccessReview
Caspase-5: Structure, Pro-Inflammatory Activity and Evolution
by
Leopold Eckhart and Heinz Fischer
Biomolecules 2024, 14(5), 520; https://doi.org/10.3390/biom14050520 - 26 Apr 2024
Abstract
Caspase-5 is a protease that induces inflammation in response to lipopolysaccharide (LPS), a component of the cell envelope of Gram-negative bacteria. The expression level of the CASP5 gene is very low in the basal state, but strongly increases in the presence of LPS.
[...] Read more.
Caspase-5 is a protease that induces inflammation in response to lipopolysaccharide (LPS), a component of the cell envelope of Gram-negative bacteria. The expression level of the CASP5 gene is very low in the basal state, but strongly increases in the presence of LPS. Intracellular LPS binds to the caspase activation and recruitment domain (CARD) of caspase-5, leading to the formation of a non-canonical inflammasome. Subsequently, the catalytic domain of caspase-5 cleaves gasdermin D and thereby facilitates the formation of cell membrane pores through which pro-inflammatory cytokines of the interleukin-1 family are released. Caspase-4 is also able to form a non-canonical inflammasome upon binding to LPS, but its expression is less dependent on LPS than the expression of caspase-5. Caspase-4 and caspase-5 have evolved via the duplication of a single ancestral gene in a subclade of primates, including humans. Notably, the main biomedical model species, the mouse, has only one ortholog, namely caspase-11. Here, we review the structural features and the mechanisms of regulation that are important for the pro-inflammatory roles of caspase-5. We summarize the interspecies differences and the evolution of pro-inflammatory caspases in mammals and discuss the potential roles of caspase-5 in the defense against Gram-negative bacteria and in sepsis.
Full article
(This article belongs to the Section Cellular Biochemistry)
►▼
Show Figures
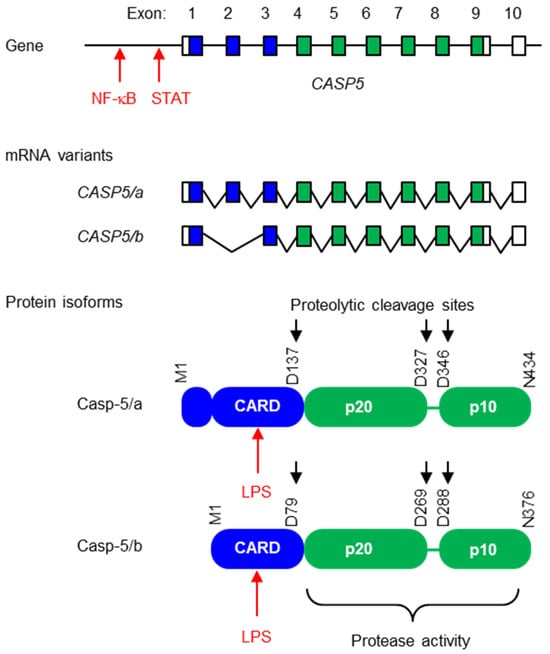
Figure 1
Open AccessArticle
Modeling Radiation-Induced Epithelial Cell Injury in Murine Three-Dimensional Esophageal Organoids
by
Latisha Carswell, Deepa M. Sridharan, Lung-Chang Chien, Wataru Hirose, Véronique Giroux, Hiroshi Nakagawa and Janice M. Pluth
Biomolecules 2024, 14(5), 519; https://doi.org/10.3390/biom14050519 (registering DOI) - 25 Apr 2024
Abstract
Esophageal squamous cell carcinoma (ESCC) is a deadly consequence of radiation exposure to the esophagus. ESCC arises from esophageal epithelial cells that undergo malignant transformation and features a perturbed squamous cell differentiation program. Understanding the dose- and radiation quality-dependence of the esophageal epithelium
[...] Read more.
Esophageal squamous cell carcinoma (ESCC) is a deadly consequence of radiation exposure to the esophagus. ESCC arises from esophageal epithelial cells that undergo malignant transformation and features a perturbed squamous cell differentiation program. Understanding the dose- and radiation quality-dependence of the esophageal epithelium response to radiation may provide insights into the ability of radiation to promote ESCC. We have explored factors that may play a role in esophageal epithelial radiosensitivity and their potential relationship to ESCC risk. We have utilized a murine three-dimensional (3D) organoid model that recapitulates the morphology and functions of the stratified squamous epithelium of the esophagus to study persistent dose- and radiation quality-dependent changes. Interestingly, although high-linear energy transfer (LET) Fe ion exposure induced a more intense and persistent alteration of squamous differentiation and 53BP1 DNA damage foci levels as compared to Cs, the MAPK/SAPK stress pathway signaling showed similar altered levels for most phospho-proteins with both radiation qualities. In addition, the lower dose of high-LET exposure also revealed nearly the same degree of morphological changes, even though only ~36% of the cells were predicted to be hit at the lower 0.1 Gy dose, suggesting that a bystander effect may be induced. Although p38 and ERK/MAPK revealed the highest levels following high-LET exposure, the findings reveal that even a low dose (0.1 Gy) of both radiation qualities can elicit a persistent stress signaling response that may critically impact the differentiation gradient of the esophageal epithelium, providing novel insights into the pathogenesis of radiation-induced esophageal injury and early stage esophageal carcinogenesis.
Full article
(This article belongs to the Special Issue Esophageal Diseases: Molecular Basis and Therapeutic Approaches)
►▼
Show Figures
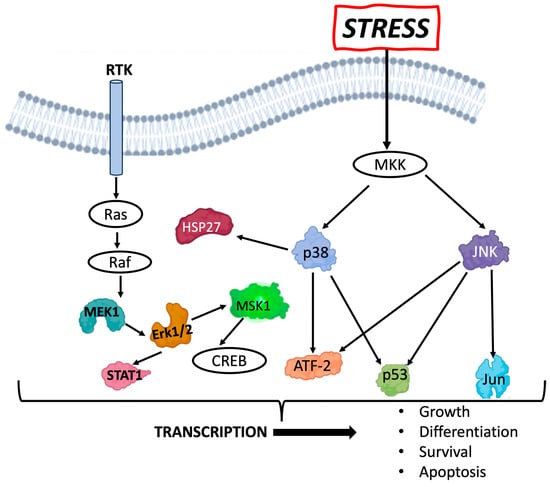
Figure 1
Open AccessFeature PaperReview
Reflections on the Origin of Coded Protein Biosynthesis
by
Juan Carlos Fontecilla-Camps
Biomolecules 2024, 14(5), 518; https://doi.org/10.3390/biom14050518 - 25 Apr 2024
Abstract
The principle of continuity posits that some central features of primordial biocatalytic mechanisms should still be present in the genetically dependent pathway of protein synthesis, a crucial step in the emergence of life. Key bimolecular reactions of this process are catalyzed by DNA-dependent
[...] Read more.
The principle of continuity posits that some central features of primordial biocatalytic mechanisms should still be present in the genetically dependent pathway of protein synthesis, a crucial step in the emergence of life. Key bimolecular reactions of this process are catalyzed by DNA-dependent RNA polymerases, aminoacyl-tRNA synthetases, and ribosomes. Remarkably, none of these biocatalysts contribute chemically active groups to their respective reactions. Instead, structural and functional studies have demonstrated that nucleotidic α-phosphate and β-d-ribosyl 2′ OH and 3′ OH groups can help their own catalysis, a process which, consequently, has been called “substrate-assisted”. Furthermore, upon binding, the substrates significantly lower the entropy of activation, exclude water from these catalysts’ active sites, and are readily positioned for a reaction. This binding mode has been described as an “entropy trap”. The combination of this effect with substrate-assisted catalysis results in reactions that are stereochemically and mechanistically simpler than the ones found in most modern enzymes. This observation is consistent with the way in which primordial catalysts could have operated; it may also explain why, thanks to their complementary reactivities, β-d-ribose and phosphate were naturally selected to be the central components of early coding polymers.
Full article
(This article belongs to the Section Enzymology)
Open AccessArticle
The Role of Cytokinins and Abscisic Acid in the Growth, Development and Virulence of the Pathogenic Fungus Stagonospora nodorum (Berk.)
by
Tatyana V. Nuzhnaya, Antonina V. Sorokan, Guzel F. Burkhanova, Igor V. Maksimov and Svetlana V. Veselova
Biomolecules 2024, 14(5), 517; https://doi.org/10.3390/biom14050517 - 25 Apr 2024
Abstract
Cytokinins (CKs) and abscisic acid (ABA) play an important role in the life of both plants and pathogenic fungi. However, the role of CKs and ABA in the regulation of fungal growth, development and virulence has not been sufficiently studied. We compared the
[...] Read more.
Cytokinins (CKs) and abscisic acid (ABA) play an important role in the life of both plants and pathogenic fungi. However, the role of CKs and ABA in the regulation of fungal growth, development and virulence has not been sufficiently studied. We compared the ability of two virulent isolates (SnB and Sn9MN-3A) and one avirulent isolate (Sn4VD) of the pathogenic fungus Stagonospora nodorum Berk. to synthesize three groups of hormones (CKs, ABA and auxins) and studied the effect of exogenous ABA and zeatin on the growth, sporulation and gene expression of necrotrophic effectors (NEs) and transcription factors (TFs) in them. Various isolates of S. nodorum synthesized different amounts of CKs, ABA and indoleacetic acid. Using exogenous ABA and zeatin, we proved that the effect of these hormones on the growth and sporulation of S. nodorum isolates can be opposite, depends on both the genotype of the isolate and on the concentration of the hormone and is carried out through the regulation of carbohydrate metabolism. ABA and zeatin regulated the expression of fungal TF and NE genes, but correlation analysis of these parameters showed that this effect depended on the genotype of the isolate. This study will contribute to our understanding of the role of the hormones ABA and CKs in the biology of the fungal pathogen S. nodorum.
Full article
(This article belongs to the Special Issue Phytohormones 2022–2023)
►▼
Show Figures
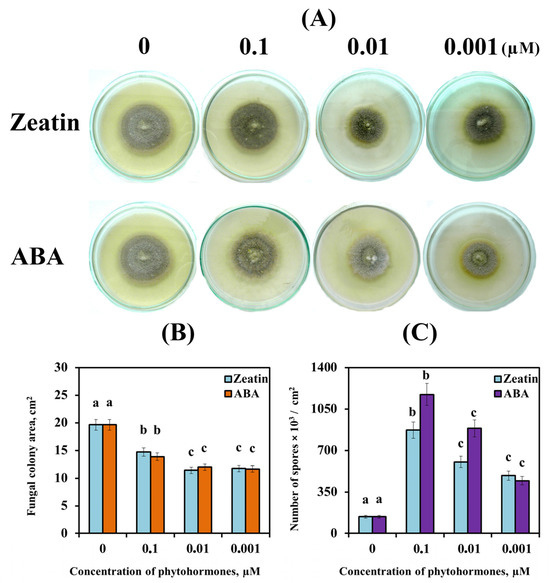
Figure 1
Open AccessArticle
GateView: A Multi-Omics Platform for Gene Feature Analysis of Virus Receptors within Human Normal Tissues and Tumors
by
Yang Sun, Zi-Liang Huang, Wen-Xin Chen, Yi-Feng Zhang, Hao-Tian Lei, Qiao-Juan Huang, Zhao-Rong Lun, Liang-Hu Qu and Ling-Ling Zheng
Biomolecules 2024, 14(5), 516; https://doi.org/10.3390/biom14050516 - 25 Apr 2024
Abstract
Viruses are obligate intracellular parasites that rely on cell surface receptor molecules to complete the first step of invading host cells. The experimental method for virus receptor screening is time-consuming, and receptor molecules have been identified for less than half of known viruses.
[...] Read more.
Viruses are obligate intracellular parasites that rely on cell surface receptor molecules to complete the first step of invading host cells. The experimental method for virus receptor screening is time-consuming, and receptor molecules have been identified for less than half of known viruses. This study collected known human viruses and their receptor molecules. Through bioinformatics analysis, common characteristics of virus receptor molecules (including sequence, expression, mutation, etc.) were obtained to study why these membrane proteins are more likely to become virus receptors. An in-depth analysis of the cataloged virus receptors revealed several noteworthy findings. Compared to other membrane proteins, human virus receptors generally exhibited higher expression levels and lower sequence conservation. These receptors were found in multiple tissues, with certain tissues and cell types displaying significantly higher expression levels. While most receptor molecules showed noticeable age-related variations in expression across different tissues, only a limited number of them exhibited gender-related differences in specific tissues. Interestingly, in contrast to normal tissues, virus receptors showed significant dysregulation in various types of tumors, particularly those associated with dsRNA and retrovirus receptors. Finally, GateView, a multi-omics platform, was established to analyze the gene features of virus receptors in human normal tissues and tumors. Serving as a valuable resource, it enables the exploration of common patterns among virus receptors and the investigation of virus tropism across different tissues, population preferences, virus pathogenicity, and oncolytic virus mechanisms.
Full article
(This article belongs to the Section Bioinformatics and Systems Biology)
►▼
Show Figures
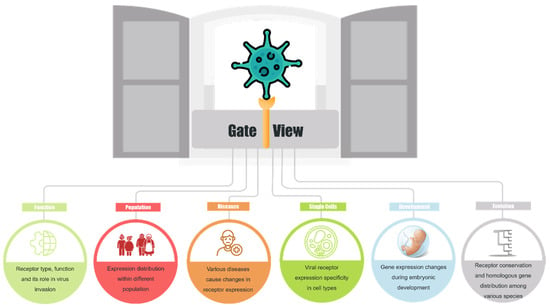
Figure 1
Open AccessReview
Fundus Autofluorescence in Posterior and Panuveitis—An Under-Estimated Imaging Technique: A Review and Case Series
by
Matthias M. Mauschitz, Markus Zeller, Pradeep Sagar, Suchitra Biswal, Gabriela Guzman, Jan H. Terheyden, Carsten H. Meyer, Frank G. Holz, Carsten Heinz, Uwe Pleyer, Robert P. Finger and Maximilian W. M. Wintergerst
Biomolecules 2024, 14(5), 515; https://doi.org/10.3390/biom14050515 - 25 Apr 2024
Abstract
Fundus autofluorescence (FAF) is a prompt and non-invasive imaging modality helpful in detecting pathological abnormalities within the retina and the choroid. This narrative review and case series provides an overview on the current application of FAF in posterior and panuveitis. The literature was
[...] Read more.
Fundus autofluorescence (FAF) is a prompt and non-invasive imaging modality helpful in detecting pathological abnormalities within the retina and the choroid. This narrative review and case series provides an overview on the current application of FAF in posterior and panuveitis. The literature was reviewed for articles on lesion characteristics on FAF of specific posterior and panuveitis entities as well as benefits and limitations of FAF for diagnosing and monitoring disease. FAF characteristics are described for non-infectious and infectious uveitis forms as well as masquerade syndromes. Dependent on the uveitis entity, FAF is of diagnostic value in detecting disease and following the clinical course. Currently available FAF modalities which differ in excitation wavelengths can provide different pathological insights depending on disease entity and activity. Further studies on the comparison of FAF modalities and their individual value for uveitis diagnosis and monitoring are warranted.
Full article
(This article belongs to the Special Issue Molecular Pathology, Diagnostics and Therapeutics of Retinal Diseases)
►▼
Show Figures
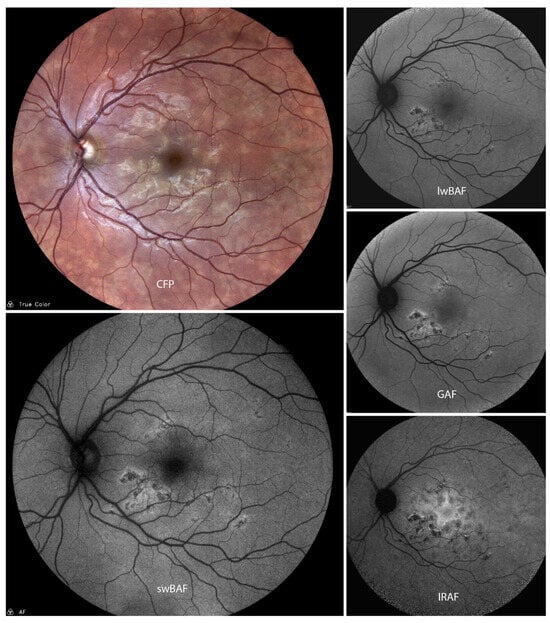
Figure 1
Open AccessReview
Novel Insights into the Links between N6-Methyladenosine and Regulated Cell Death in Musculoskeletal Diseases
by
Juanjuan Han, Cuijing Wang, Haolin Yang, Jiayi Luo, Xiaoyi Zhang and Xin-An Zhang
Biomolecules 2024, 14(5), 514; https://doi.org/10.3390/biom14050514 (registering DOI) - 24 Apr 2024
Abstract
Musculoskeletal diseases (MSDs), including osteoarthritis (OA), osteosarcoma (OS), multiple myeloma (MM), intervertebral disc degeneration (IDD), osteoporosis (OP), and rheumatoid arthritis (RA), present noteworthy obstacles associated with pain, disability, and impaired quality of life on a global scale. In recent years, it has become
[...] Read more.
Musculoskeletal diseases (MSDs), including osteoarthritis (OA), osteosarcoma (OS), multiple myeloma (MM), intervertebral disc degeneration (IDD), osteoporosis (OP), and rheumatoid arthritis (RA), present noteworthy obstacles associated with pain, disability, and impaired quality of life on a global scale. In recent years, it has become increasingly apparent that N6-methyladenosine (m6A) is a key regulator in the expression of genes in a multitude of biological processes. m6A is composed of 0.1–0.4% adenylate residues, especially at the beginning of 3′-UTR near the translation stop codon. The m6A regulator can be classified into three types, namely the “writer”, “reader”, and “eraser”. Studies have shown that the epigenetic modulation of m6A influences mRNA processing, nuclear export, translation, and splicing. Regulated cell death (RCD) is the autonomous and orderly death of cells under genetic control to maintain the stability of the internal environment. Moreover, distorted RCDs are widely used to influence the course of various diseases and receiving increasing attention from researchers. In the past few years, increasing evidence has indicated that m6A can regulate gene expression and thus influence different RCD processes, which has a central role in the etiology and evolution of MSDs. The RCDs currently confirmed to be associated with m6A are autophagy-dependent cell death, apoptosis, necroptosis, pyroptosis, ferroptosis, immunogenic cell death, NETotic cell death and oxeiptosis. The m6A–RCD axis can regulate the inflammatory response in chondrocytes and the invasive and migratory of MM cells to bone remodeling capacity, thereby influencing the development of MSDs. This review gives a complete overview of the regulatory functions on the m6A–RCD axis across muscle, bone, and cartilage. In addition, we also discuss recent advances in the control of RCD by m6A-targeted factors and explore the clinical application prospects of therapies targeting the m6A–RCD in MSD prevention and treatment. These may provide new ideas and directions for understanding the pathophysiological mechanism of MSDs and the clinical prevention and treatment of these diseases.
Full article
(This article belongs to the Topic Bone-Related Diseases: From Molecular Mechanisms to Therapy Development)
►▼
Show Figures
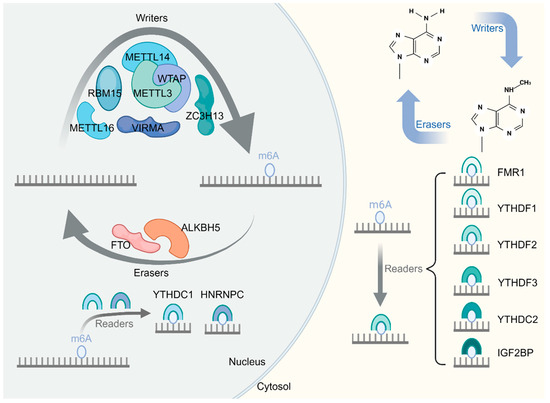
Figure 1
Open AccessReview
The Exploitation of the Glycosylation Pattern in Asthma: How We Alter Ancestral Pathways to Develop New Treatments
by
Angelika Muchowicz, Agnieszka Bartoszewicz and Zbigniew Zaslona
Biomolecules 2024, 14(5), 513; https://doi.org/10.3390/biom14050513 - 24 Apr 2024
Abstract
Asthma has reached epidemic levels, yet progress in developing specific therapies is slow. One of the main reasons for this is the fact that asthma is an umbrella term for various distinct subsets. Due to its high heterogeneity, it is difficult to establish
[...] Read more.
Asthma has reached epidemic levels, yet progress in developing specific therapies is slow. One of the main reasons for this is the fact that asthma is an umbrella term for various distinct subsets. Due to its high heterogeneity, it is difficult to establish biomarkers for each subset of asthma and to propose endotype-specific treatments. This review focuses on protein glycosylation as a process activated in asthma and ways to utilize it to develop novel biomarkers and treatments. We discuss known and relevant glycoproteins whose functions control disease development. The key role of glycoproteins in processes integral to asthma, such as inflammation, tissue remodeling, and repair, justifies our interest and research in the field of glycobiology. Altering the glycosylation states of proteins contributing to asthma can change the pathological processes that we previously failed to inhibit. Special emphasis is placed on chitotriosidase 1 (CHIT1), an enzyme capable of modifying LacNAc- and LacdiNAc-containing glycans. The expression and activity of CHIT1 are induced in human diseased lungs, and its pathological role has been demonstrated by both genetic and pharmacological approaches. We propose that studying the glycosylation pattern and enzymes involved in glycosylation in asthma can help in patient stratification and in developing personalized treatment.
Full article
(This article belongs to the Special Issue Biomarkers of Severe Asthma: From Pathogenetic Mechanisms to Clinical Application)
►▼
Show Figures
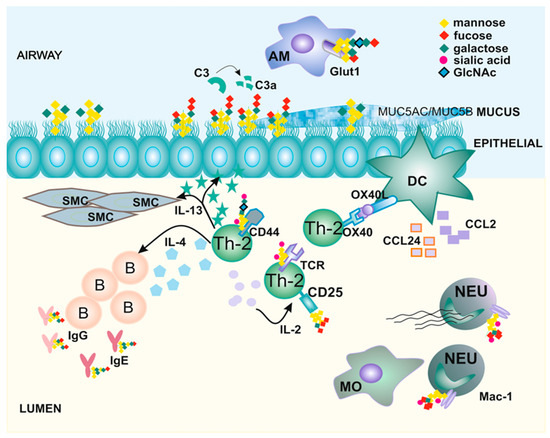
Figure 1

Journal Menu
► ▼ Journal Menu-
- Biomolecules Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Bioengineering, Biomolecules, Cancers, Diseases, Nanomaterials, Pharmaceutics
Dynamic Nano-Biomaterials in Tissue Regeneration and Cancer Therapies
Topic Editors: Ramar Thangam, Heemin Kang, Bibin G. Anand, Ramachandran Vijayan, Venugopal KrishnanDeadline: 15 May 2024
Topic in
Analytica, Biomolecules, Metabolites, Molecules, Separations
Advances in Separation Methods for Metabolomics and Lipidomics
Topic Editors: Eduardo Sommella, Giulia Mazzoccanti, Emanuela SalviatiDeadline: 31 May 2024
Topic in
BioChem, Biomedicines, Biomolecules, IJMS, Metabolites, Molecules
Natural Products in Prevention and Therapy of Metabolic Syndrome
Topic Editors: Jianbo Wan, Ligen LinDeadline: 30 June 2024
Topic in
Biomolecules, Cancers, Current Oncology, JCM, Onco
Extracellular Vesicles in Cancer Diagnosis and Treatment
Topic Editors: Shenglin Huang, Linlin Guo, Jialei WangDeadline: 20 July 2024

Conferences
Special Issues
Special Issue in
Biomolecules
PPARs as Key Regulators in Different Diseases
Guest Editors: Roberta Montanari, Davide CapelliDeadline: 30 April 2024
Special Issue in
Biomolecules
Molecular Pathology, Diagnostics and Therapeutics of Retinal Diseases
Guest Editor: Antonio FerrerasDeadline: 15 May 2024
Special Issue in
Biomolecules
Neuroanatomy as an Implement for the Study of CNS Function and Dysfunction
Guest Editors: Anastasia Tsingotjidou, Chryssa BekiariDeadline: 31 May 2024
Special Issue in
Biomolecules
Role of Neuron-Glia and Glia-Glia Communications in the Brain Function: Recent Advances in the Central Nervous System Cellular Crosstalk and Signaling
Guest Editor: Nima SanadgolDeadline: 15 June 2024
Topical Collections
Topical Collection in
Biomolecules
Feature Papers in Molecular Genetics
Collection Editor: Jürg Bähler
Topical Collection in
Biomolecules
Feature Papers in Bioinformatics and Systems Biology Section
Collection Editor: Lukasz Kurgan
Topical Collection in
Biomolecules
Molecular Mechanisms of Obesity, Diabetes, Inflammation and Aging
Collection Editors: Yuxiang Sun, Susanne Talcott