Journal Description
Viruses
Viruses
is a peer-reviewed, open access journal of virology, published monthly online by MDPI. The American Society for Virology (ASV), Spanish Society for Virology (SEV), Canadian Society for Virology (CSV), Italian Society for Virology (SIV-ISV), Australasian Virology Society (AVS) and others are affiliated with Viruses and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, MEDLINE, PMC, Embase, PubAg, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Virology) / CiteScore - Q1 (Infectious Diseases)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 13.8 days after submission; acceptance to publication is undertaken in 2.5 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Companion journals for Viruses include: COVID and Zoonotic Diseases.
Impact Factor:
4.7 (2022);
5-Year Impact Factor:
4.8 (2022)
Latest Articles
Virus Quasispecies Rarefaction: Subsampling with or without Replacement?
Viruses 2024, 16(5), 710; https://doi.org/10.3390/v16050710 (registering DOI) - 29 Apr 2024
Abstract
In quasispecies diversity studies, the comparison of two samples of varying sizes is a common necessity. However, the sensitivity of certain diversity indices to sample size variations poses a challenge. To address this issue, rarefaction emerges as a crucial tool, serving to normalize
[...] Read more.
In quasispecies diversity studies, the comparison of two samples of varying sizes is a common necessity. However, the sensitivity of certain diversity indices to sample size variations poses a challenge. To address this issue, rarefaction emerges as a crucial tool, serving to normalize and create fairly comparable samples. This study emphasizes the imperative nature of sample size normalization in quasispecies diversity studies using next-generation sequencing (NGS) data. We present a thorough examination of resampling schemes using various simple hypothetical cases of quasispecies showing different quasispecies structures in the sense of haplotype genomic composition, offering a comprehensive understanding of their implications in general cases. Despite the big numbers implied in this sort of study, often involving coverages exceeding 100,000 reads per sample and amplicon, the rarefaction process for normalization should be performed with repeated resampling without replacement, especially when rare haplotypes constitute a significant fraction of interest. However, it is noteworthy that different diversity indices exhibit distinct sensitivities to sample size. Consequently, some diversity indicators may be compared directly without normalization, or instead may be resampled safely with replacement.
Full article
(This article belongs to the Section General Virology)
►
Show Figures
Open AccessCase Report
COVID-19 in Lung Transplant Recipients: A Report on 10 Recent Cases
by
Lea Reemann, Nikolaus Kneidinger, Bernd Sczepanski and Andreas Rembert Koczulla
Viruses 2024, 16(5), 709; https://doi.org/10.3390/v16050709 (registering DOI) - 29 Apr 2024
Abstract
Due to immunosuppression, transplant recipients are at higher risk of infections with SARS-CoV-2 and worse clinical outcomes than immunocompetent hosts. Furthermore, lung transplant patients represent a special group among solid organ recipients, since pneumonia is the main manifestation of COVID-19. However, data on
[...] Read more.
Due to immunosuppression, transplant recipients are at higher risk of infections with SARS-CoV-2 and worse clinical outcomes than immunocompetent hosts. Furthermore, lung transplant patients represent a special group among solid organ recipients, since pneumonia is the main manifestation of COVID-19. However, data on the course of disease and the changes in morbidity and mortality during the course of the pandemic are limited. In our pulmonary rehabilitation clinic, we treat patients shortly after lung transplant as well as long-term transplant patients. Over the last almost 4 years of pandemic, we witnessed several COVID-19 infections in lung transplant patients in our clinic as well as patients who acquired an infection beforehand. In this paper, we aim at retrospectively describing a series of recent COVID-19 cases in our clinic, looking at the clinical course of disease and outcomes in lung transplant patients.
Full article
(This article belongs to the Section Coronaviruses)
Open AccessArticle
Development of a Cell Culture Model for Inducible SARS-CoV-2 Replication
by
Xiaoyan Wang, Yuanfei Zhu, Qiong Wu, Nan Jiang, Youhua Xie and Qiang Deng
Viruses 2024, 16(5), 708; https://doi.org/10.3390/v16050708 (registering DOI) - 29 Apr 2024
Abstract
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) induces direct cytopathic effects, complicating the establishment of low-cytotoxicity cell culture models for studying its replication. We initially developed a DNA vector-based replicon system utilizing the CMV promoter to generate a recombinant viral genome bearing reporter
[...] Read more.
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) induces direct cytopathic effects, complicating the establishment of low-cytotoxicity cell culture models for studying its replication. We initially developed a DNA vector-based replicon system utilizing the CMV promoter to generate a recombinant viral genome bearing reporter genes. However, this system frequently resulted in drug resistance and cytotoxicity, impeding model establishment. Herein, we present a novel cell culture model with SARS-CoV-2 replication induced by Cre/LoxP-mediated DNA recombination. An engineered SARS-CoV-2 transcription unit was subcloned into a bacterial artificial chromosome (BAC) vector. To enhance biosafety, the viral spike protein gene was deleted, and the nucleocapsid gene was replaced with a reporter gene. An exogenous sequence was inserted within NSP1 as a modulatory cassette that is removable after Cre/LoxP-mediated DNA recombination and subsequent RNA splicing. Using the PiggyBac transposon strategy, the transcription unit was integrated into host cell chromatin, yielding a stable cell line capable of inducing recombinant SARS-CoV-2 RNA replication. The model exhibited sensitivity to the potential antivirals forsythoside A and verteporfin. An innovative inducible SARS-CoV-2 replicon cell model was introduced to further explore the replication and pathogenesis of the virus and facilitate screening and assessment of anti-SARS-CoV-2 therapeutics.
Full article
(This article belongs to the Special Issue Novel Antiviral Agents: Synthesis, Molecular Modelling Studies and Biological Investigation, 2nd Edition)
Open AccessReview
The Adaptive Immune Response in Hepatitis B Virus-Associated Hepatocellular Carcinoma Is Characterized by Dysfunctional and Exhausted HBV-Specific T Cells
by
Malene Broholm, Anne-Sofie Mathiasen, Ása Didriksen Apol and Nina Weis
Viruses 2024, 16(5), 707; https://doi.org/10.3390/v16050707 (registering DOI) - 29 Apr 2024
Abstract
This systematic review investigates the immunosuppressive environment in HBV-associated hepatocellular carcinoma (HCC), characterized by dysfunctional and exhausted HBV-specific T cells alongside an increased infiltration of HBV-specific CD4+ T cells, particularly regulatory T cells (Tregs). Heightened expression of checkpoint inhibitors, notably PD-1, is linked
[...] Read more.
This systematic review investigates the immunosuppressive environment in HBV-associated hepatocellular carcinoma (HCC), characterized by dysfunctional and exhausted HBV-specific T cells alongside an increased infiltration of HBV-specific CD4+ T cells, particularly regulatory T cells (Tregs). Heightened expression of checkpoint inhibitors, notably PD-1, is linked with disease progression and recurrence, indicating its potential as both a prognostic indicator and a target for immunotherapy. Nevertheless, using PD-1 inhibitors has shown limited effectiveness. In a future perspective, understanding the intricate interplay between innate and adaptive immune responses holds promise for pinpointing predictive biomarkers and crafting novel treatment approaches for HBV-associated HCC.
Full article
(This article belongs to the Section Human Virology and Viral Diseases)
Open AccessArticle
Additional Insertion of gC Gene Triggers Better Immune Efficacy of TK/gI/gE-Deleted Pseudorabies Virus in Mice
by
Zhuoyun Wu, Jiahuan Deng, Meijing Chen, Peiqi Lu, Zhibin Yan, Xiaoyan Wu, Qiuyun Ji, Huiying Fan, Yongwen Luo and Chunmei Ju
Viruses 2024, 16(5), 706; https://doi.org/10.3390/v16050706 (registering DOI) - 29 Apr 2024
Abstract
In recent years, pseudorabies virus (PRV) variants have resulted in an epidemic in swine herds and huge economic losses in China. Therefore, it is essential to develop an efficacious vaccine against the spread of PRV variants. Here, the triple-gene-deletion virus and the triple-gene-deletion
[...] Read more.
In recent years, pseudorabies virus (PRV) variants have resulted in an epidemic in swine herds and huge economic losses in China. Therefore, it is essential to develop an efficacious vaccine against the spread of PRV variants. Here, the triple-gene-deletion virus and the triple-gene-deletion plus gC virus were constructed by homologous recombination (HR). And then, their growth capacity, proliferation ability, and immune efficacy were evaluated. The results showed that the growth kinetics of the recombinant viruses were similar to those of the parental strain PRV-AH. Compared with the triple-gene-deletion virus group, the more dominant level of neutralizing antibody (NA) can be induced in the triple-gene-deletion plus gC virus group with the same 106.0 TCID50 dose after 4 and 6 weeks post-initial immunization (PII) (p < 0.0001). In addition, the antibody titers in mice immunized with the triple-gene-deletion plus gC virus were significantly higher than those immunized with triple-gene deletion virus with the same 105.0 TCID50 dose after 6 weeks PII (p < 0.001). More importantly, in the triple-gene-deletion plus gC virus group with 105.0 TCID50, the level of NA was close to that in the triple-gene deletion virus group with 106.0 TCID50 at 6 weeks PII. Meanwhile, the cytokines IL-4 and IFN-γ in sera were tested by enzyme-linked immunosorbent assay (ELISA) in each group. The highest level of IL-4 or IFN-γ was also elicited in the triple-gene deletion plus gC virus group at a dose of 106.0 TCID50. After challenge with PRV-AH, the survival rates of the triple-gene deletion plus gC virus immunized groups were higher than those of other groups. In immunized groups with 105.0 TCID50, the survival rate shows a significant difference between the triple-gene deletion plus gC virus group (75%, 6/8) and the triple-gene deletion virus group (12.5%, 1/8). In general, the immune efficacy of the PRV TK/gI/gE-deleted virus can be increased with additional gC insertion in mice, which has potential for developing an attenuated vaccine candidate for PRV control.
Full article
(This article belongs to the Section Animal Viruses)
►▼
Show Figures
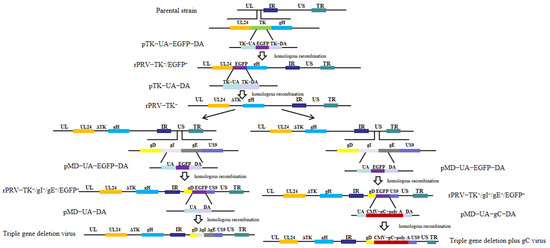
Figure 1
Open AccessArticle
Determination and Characterization of Novel Papillomavirus and Parvovirus Associated with Mass Mortality of Chinese Tongue Sole (Cynoglossus semilaevis) in China
by
Shuxia Xue, Xinrui Liu, Yuru Liu, Chang Lu, Lei Jia, Yanguang Yu, Houfu Liu, Siyu Yang, Zhu Zeng, Hui Li, Jiatong Qin, Yuxuan Wang and Jinsheng Sun
Viruses 2024, 16(5), 705; https://doi.org/10.3390/v16050705 (registering DOI) - 29 Apr 2024
Abstract
A massive mortality event concerning farmed Chinese tongue soles occurred in Tianjin, China, and the causative agent remains unknown. Here, a novel Cynoglossus semilaevis papillomavirus (CsPaV) and parvovirus (CsPV) were simultaneously isolated and identified from diseased fish via electron microscopy, virus isolation, genome
[...] Read more.
A massive mortality event concerning farmed Chinese tongue soles occurred in Tianjin, China, and the causative agent remains unknown. Here, a novel Cynoglossus semilaevis papillomavirus (CsPaV) and parvovirus (CsPV) were simultaneously isolated and identified from diseased fish via electron microscopy, virus isolation, genome sequencing, experimental challenges, and fluorescence in situ hybridization (FISH). Electron microscopy showed large numbers of virus particles present in the tissues of diseased fish. Viruses that were isolated and propagated in flounder gill cells (FG) induced typical cytopathic effects (CPE). The cumulative mortality of fish given intraperitoneal injections reached 100% at 7 dpi. The complete genomes of CsPaV and CsPV comprised 5939 bp and 3663 bp, respectively, and the genomes shared no nucleotide sequence similarities with other viruses. Phylogenetic analysis based on the L1 and NS1 protein sequences revealed that CsPaV and CsPV were novel members of the Papillomaviridae and Parvoviridae families. The FISH results showed positive signals in the spleen tissues of infected fish, and both viruses could co-infect single cells. This study represents the first report where novel papillomavirus and parvovirus are identified in farmed marine cultured fish, and it provides a basis for further studies on the prevention and treatment of emerging viral diseases.
Full article
(This article belongs to the Special Issue Animal Virus Discovery and Genetic Diversity)
►▼
Show Figures
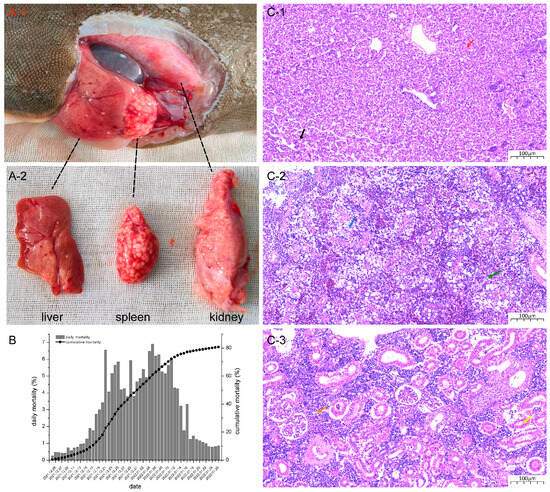
Figure 1
Open AccessArticle
Epidemiology of Respiratory Syncytial Virus Hospitalizations in Poland: An Analysis from 2015 to 2023 Covering the Entire Polish Population of Children Aged under Five Years
by
Jan Mazela, Teresa Jackowska, Marcin Czech, Ewa Helwich, Oliver Martyn, Pawel Aleksiejuk, Anna Smaga, Joanna Glazewska and Jacek Wysocki
Viruses 2024, 16(5), 704; https://doi.org/10.3390/v16050704 (registering DOI) - 29 Apr 2024
Abstract
Background: Respiratory syncytial virus (RSV) is an important cause of childhood hospitalizations. The aim of the study was to estimate the rates of RSV-related hospitalizations in children aged less than 5 years in Poland. Methods: This retrospective observational cohort study was based on
[...] Read more.
Background: Respiratory syncytial virus (RSV) is an important cause of childhood hospitalizations. The aim of the study was to estimate the rates of RSV-related hospitalizations in children aged less than 5 years in Poland. Methods: This retrospective observational cohort study was based on data obtained from the National Health Fund in Poland regarding all acute respiratory tract infections and RSV-coded admissions of children (age <5 years) to public hospitals between July 2015 and June 2023. Patients were stratified based on the following age groups: 0–1 month, 2–3 months, 4–6 months, 7–12 months, 13–24 months, and 25–60 months. Results: The number of RSV-related hospitalizations increased every season, both before and through the ending phase of the coronavirus disease 2019 (COVID-19) pandemic. The COVID-19 pandemic was associated with a shift in the seasonality pattern of RSV infection. Hospitalization rates per 1000 inhabitants were the highest for children aged 0–12 months, reaching 47.3 in the 2022/23 season. Within this group, the highest hospitalization rate was observed for children aged 2–3 months—94.9 in the 2022/23 season. During the ending phase of the COVID-19 pandemic, the observed increase in admission rates was 2-, 4-, and 5-fold the pre-COVID rate for children aged <12 months, 12–24 months, and 25–60 months, respectively. Conclusions: In Poland, RSV infections cause a significant burden in hospitalized children aged less than 5 years. RSV-related hospitalizations were most frequent in children aged less than 1 year. The COVID-19 pandemic was associated with a shift in the seasonality pattern of RSV infections. After the pandemic, more RSV-related hospitalizations were observed in older children (aged 13 months and older) vs. the pre-pandemic phase.
Full article
(This article belongs to the Section Human Virology and Viral Diseases)
Open AccessReview
Winged Threat on the Offensive: A Literature Review Due to the First Identification of Aedes japonicus in Poland
by
Marcin Gierek, Gabriela Ochała-Gierek, Andrzej Józef Woźnica, Grzegorz Zaleśny, Alicja Jarosz and Paweł Niemiec
Viruses 2024, 16(5), 703; https://doi.org/10.3390/v16050703 (registering DOI) - 29 Apr 2024
Abstract
Genetic studies preceded by the observation of an unknown mosquito species in Mikołów (Poland) confirmed that it belongs to a new invasive species in Polish fauna, Aedes japonicus (Theobald, 1901), a known vector for numerous infectious diseases. Ae. japonicus is expanding its geographical
[...] Read more.
Genetic studies preceded by the observation of an unknown mosquito species in Mikołów (Poland) confirmed that it belongs to a new invasive species in Polish fauna, Aedes japonicus (Theobald, 1901), a known vector for numerous infectious diseases. Ae. japonicus is expanding its geographical presence, raising concerns about potential disease transmission given its vector competence for chikungunya virus, dengue virus, West Nile virus, and Zika virus. This first genetically confirmed identification of Ae. japonicus in Poland initiates a comprehensive review of the literature on Ae. japonicus, its biology and ecology, and the viral infections transmitted by this species. This paper also presents the circumstances of the observation of Ae. japonicus in Poland and a methodology for identifying this species.
Full article
(This article belongs to the Section Human Virology and Viral Diseases)
►▼
Show Figures
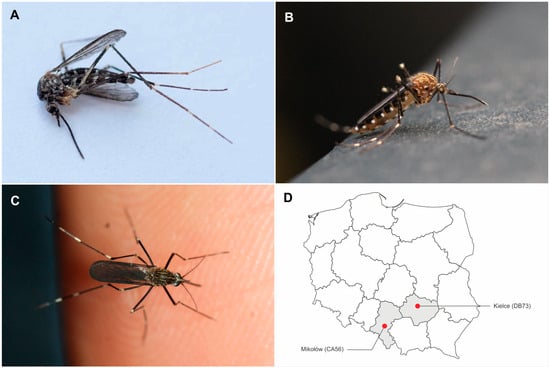

Figure 1
Open AccessArticle
HACD3 Prevents PB1 from Autophagic Degradation to Facilitate the Replication of Influenza A Virus
by
Qibing Li, Li Jiang, Yihan Wang, Xuwei Liu, Bo Wang, Zhibo Shan, Yi-Han Wang, Yuqin Wang, Hualan Chen and Chengjun Li
Viruses 2024, 16(5), 702; https://doi.org/10.3390/v16050702 (registering DOI) - 29 Apr 2024
Abstract
Influenza A virus (IAV) continues to pose serious threats to the global animal industry and public health security. Identification of critical host factors engaged in the life cycle of IAV and elucidation of the underlying mechanisms of their action are particularly important for
[...] Read more.
Influenza A virus (IAV) continues to pose serious threats to the global animal industry and public health security. Identification of critical host factors engaged in the life cycle of IAV and elucidation of the underlying mechanisms of their action are particularly important for the discovery of potential new targets for the development of anti-influenza drugs. Herein, we identified Hydroxyacyl-CoA Dehydratase 3 (HACD3) as a new host factor that supports the replication of IAV. Downregulating the expression of HACD3 reduced the level of viral PB1 protein in IAV-infected cells and in cells that were transiently transfected to express PB1. Silencing HACD3 expression had no effect on the level of PB1 mRNA but could promote the lysosome-mediated autophagic degradation of PB1 protein. Further investigation revealed that HACD3 interacted with PB1 and selective autophagic receptor SQSTM1/p62, and HACD3 competed with SQSTM1/p62 for the interaction with PB1, which prevented PB1 from SQSTM1/p62-mediated autophagic degradation. Collectively, these findings establish that HACD3 plays a positive regulatory role in IAV replication by stabilizing the viral PB1 protein.
Full article
(This article belongs to the Special Issue Host–Viral Protein Interactions and Post-translational Modifications in Viral Infections)
►▼
Show Figures
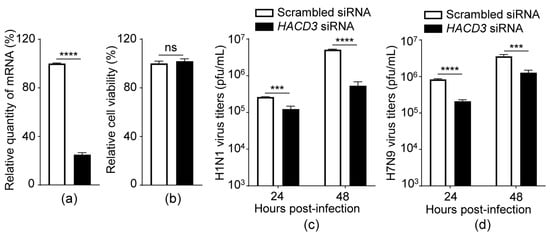
Figure 1
Open AccessArticle
Two Lineages of Papillomaviruses Identified from Caracals (Caracal caracal) in South Africa
by
Simona Kraberger, Laurel E. K. Serieys, Gabriella R. M. Leighton, Matthew D. De Koch, John S. Munday, Jacqueline M. Bishop and Arvind Varsani
Viruses 2024, 16(5), 701; https://doi.org/10.3390/v16050701 (registering DOI) - 29 Apr 2024
Abstract
Papillomaviruses (PV) infect epithelial cells and can cause hyperplastic or neoplastic lesions. In felids, most described PVs are from domestic cats (Felis catus; n = 7 types), with one type identified in each of the five wild felid species studied to
[...] Read more.
Papillomaviruses (PV) infect epithelial cells and can cause hyperplastic or neoplastic lesions. In felids, most described PVs are from domestic cats (Felis catus; n = 7 types), with one type identified in each of the five wild felid species studied to date (Panthera uncia, Puma concolor, Leopardus wiedii, Panthera leo persica and Lynx rufus). PVs from domestic cats are highly diverse and are currently classified into three genera (Lambdapapillomavirus, Dyothetapapillomavirus, and Taupapillomavirus), whereas those from wild felids, although diverse, are all classified into the Lambdapapillomavirus genus. In this study, we used a metagenomic approach to identify ten novel PV genomes from rectal swabs of five deceased caracals (Caracal caracal) living in the greater Cape Town area, South Africa. These are the first PVs to be described from caracals, and represent six new PV types, i.e., Caracal caracal papillomavirus (CcarPV) 1–6. These CcarPV fall into two phylogenetically distinct genera: Lambdapapillomavirus, and Treisetapapillomavirus. Two or more PV types were identified in a single individual for three of the five caracals, and four caracals shared at least one of the same PV types with another caracal. This study broadens our understanding of wild felid PVs and provides evidence that there may be several wild felid PV lineages.
Full article
(This article belongs to the Special Issue Animal Virus Discovery and Genetic Diversity)
►▼
Show Figures
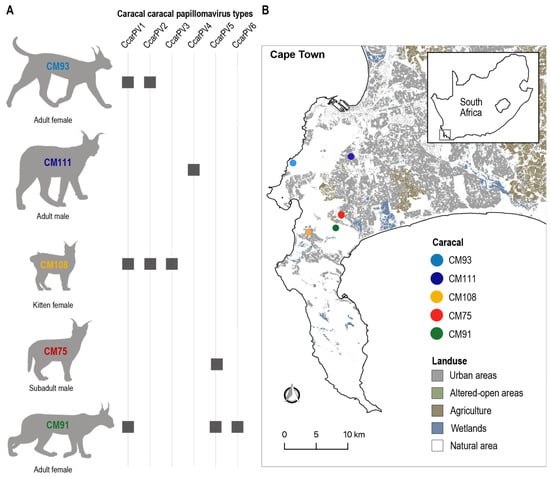
Figure 1
Open AccessBrief Report
Correlation between the Cycle Threshold Values in Detection of Severe Fever with Thrombocytopenia Syndrome Virus Using PowerChekTM SFTSV Real-Time PCR Kit and Viral Load: Prognostic Implications
by
Misun Kim, Sang Taek Heo, Hee Cheol Kim, Myeong Jin Kang, Sora Kim, Keun Hwa Lee and Jeong Rae Yoo
Viruses 2024, 16(5), 700; https://doi.org/10.3390/v16050700 (registering DOI) - 29 Apr 2024
Abstract
Background: This study aimed to analyze the correlation between the cycle threshold (Ct) values of severe fever with thrombocytopenia syndrome (SFTS) virus small (S) and middle (M) segments and the SFTS viral load, aiming to estimate the initial viral load and predict prognosis
[...] Read more.
Background: This study aimed to analyze the correlation between the cycle threshold (Ct) values of severe fever with thrombocytopenia syndrome (SFTS) virus small (S) and middle (M) segments and the SFTS viral load, aiming to estimate the initial viral load and predict prognosis in the early clinical course. Method: A retrospective study was conducted with confirmed SFTS patients at Jeju National University Hospital (2016–2022). Patients were categorized into non-fatal and fatal groups. Results: This study included 49 patients with confirmed SFTS (non-fatal group, n = 42; fatal group, n = 7). A significant negative correlation (−0.783) was observed between the log SFTS viral load and Ct values (p < 0.001). This negative correlation was notably stronger in the fatal group (correlation coefficient −0.940) than in the non-fatal group (correlation coefficient −0.345). Conclusion: In this study, we established a correlation between SFTS viral load and Ct values for estimating the initial viral load and early predicting prognosis. These results are expected to offer valuable insights for SFTS patient treatment and prognosis prediction.
Full article
(This article belongs to the Special Issue Severe Fever with Thrombocytopenia Syndrome Virus 3.0)
►▼
Show Figures
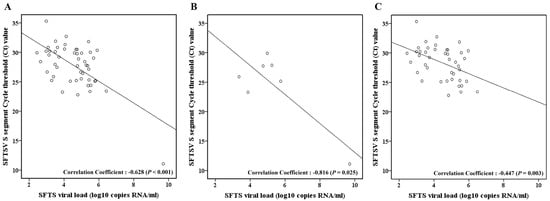
Figure 1
Open AccessCommunication
Single Amino Acid Substitution in the Matrix Protein of Rabies Virus Is Associated with Neurovirulence in Mice
by
Michiko Harada, Aya Matsuu, Yoshihiro Kaku, Akiko Okutani, Yusuke Inoue, Guillermo Posadas-Herrera, Satoshi Inoue and Ken Maeda
Viruses 2024, 16(5), 699; https://doi.org/10.3390/v16050699 (registering DOI) - 28 Apr 2024
Abstract
Rabies is a fatal encephalitic infectious disease caused by the rabies virus (RABV). RABV is highly neurotropic and replicates in neuronal cell lines in vitro. The RABV fixed strain, HEP-Flury, was produced via passaging in primary chicken embryonic fibroblast cells. HEP-Flury showed
[...] Read more.
Rabies is a fatal encephalitic infectious disease caused by the rabies virus (RABV). RABV is highly neurotropic and replicates in neuronal cell lines in vitro. The RABV fixed strain, HEP-Flury, was produced via passaging in primary chicken embryonic fibroblast cells. HEP-Flury showed rapid adaptation when propagated in mouse neuroblastoma (MNA) cells. In this study, we compared the growth of our previously constructed recombinant HEP (rHEP) strain—based on the sequence of the HEP (HEP-Flury) strain—with that of the original HEP strain. The original HEP strain exhibited higher titer than rHEP and a single substitution at position 80 in the matrix (M) protein M(D80N) after incubation in MNA cells, which was absent in rHEP. In vivo, intracerebral inoculation of the rHEP-M(D80N) strain with this substitution resulted in enhanced viral growth in the mouse brain and a significant loss of body weight in the adult mice. The number of viral antigen-positive cells in the brains of adult mice inoculated with the rHEP-M(D80N) strain was significantly higher than that with the rHEP strain at 5 days post-inoculation. Our findings demonstrate that a single amino acid substitution in the M protein M(D80N) is associated with neurovirulence in mice owing to adaptation to mouse neuronal cells.
Full article
(This article belongs to the Special Issue The World of Rhabdoviruses)
►▼
Show Figures
Figure 1
Open AccessArticle
Genetic Diversity and Detection of Respiratory Viruses Excluding SARS-CoV-2 during the COVID-19 Pandemic in Gabon, 2020–2021
by
Georgelin Nguema Ondo, Yuri Ushijima, Haruka Abe, Saïdou Mahmoudou, Rodrigue Bikangui, Anne Marie Nkoma, Marien Juliet Veraldy Magossou Mbadinga, Ayong More, Maradona Daouda Agbanrin, Christelle M. Pemba, Romuald Beh Mba, Adegnika Ayola Akim, Bertrand Lell and Jiro Yasuda
Viruses 2024, 16(5), 698; https://doi.org/10.3390/v16050698 (registering DOI) - 28 Apr 2024
Abstract
Acute respiratory infections are a major global burden in resource-limited countries, including countries in Africa. Although COVID-19 has been well studied since the pandemic emerged in Gabon, Central Africa, less attention has been paid to other respiratory viral diseases, and very little data
[...] Read more.
Acute respiratory infections are a major global burden in resource-limited countries, including countries in Africa. Although COVID-19 has been well studied since the pandemic emerged in Gabon, Central Africa, less attention has been paid to other respiratory viral diseases, and very little data are available. Herein, we provide the first data on the genetic diversity and detection of 18 major respiratory viruses in Gabon during the COVID-19 pandemic. Of 582 nasopharyngeal swab specimens collected from March 2020 to July 2021, which were SARS-CoV-2 negative, 156 were positive (26%) for the following viruses: enterovirus (20.3%), human rhinovirus (HRV) (4.6%), human coronavirus OC43 (1.2%), human adenovirus (0.9%), human metapneumovirus (hMPV) (0.5%), influenza A virus (IAV) (0.3%), and human parainfluenza viruses (0.5%). To determine the genetic diversity and transmission route of the viruses, phylogenetic analyses were performed using genome sequences of the detected viruses. The IAV strain detected in this study was genetically similar to strains isolated in the USA, whereas the hMPV strain belonging to the A2b subtype formed a cluster with Kenyan strains. This study provides the first complete genomic sequences of HRV, IAV, and hMPV detected in Gabon, and provides insight into the circulation of respiratory viruses in the country.
Full article
(This article belongs to the Special Issue The Influence of COVID-19 Pandemic on the Epidemiology of Other Human Viral Infections)
►▼
Show Figures
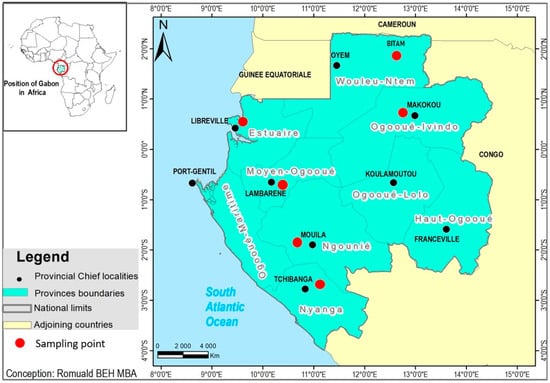
Figure 1
Open AccessReview
SARS-CoV-2 Omicron: Viral Evolution, Immune Evasion, and Alternative Durable Therapeutic Strategies
by
Hailong Guo, Sha Ha, Jason W. Botten, Kai Xu, Ningyan Zhang, Zhiqiang An, William R. Strohl, John W. Shiver and Tong-Ming Fu
Viruses 2024, 16(5), 697; https://doi.org/10.3390/v16050697 (registering DOI) - 28 Apr 2024
Abstract
Since the SARS-CoV-2 Omicron virus has gained dominance worldwide, its continual evolution with unpredictable mutations and patterns has revoked all authorized immunotherapeutics. Rapid viral evolution has also necessitated several rounds of vaccine updates in order to provide adequate immune protection. It remains imperative
[...] Read more.
Since the SARS-CoV-2 Omicron virus has gained dominance worldwide, its continual evolution with unpredictable mutations and patterns has revoked all authorized immunotherapeutics. Rapid viral evolution has also necessitated several rounds of vaccine updates in order to provide adequate immune protection. It remains imperative to understand how Omicron evolves into different subvariants and causes immune escape as this could help reevaluate the current intervention strategies mostly implemented in the clinics as emergency measures to counter the pandemic and, importantly, develop new solutions. Here, we provide a review focusing on the major events of Omicron viral evolution, including the features of spike mutation that lead to immune evasion against monoclonal antibody (mAb) therapy and vaccination, and suggest alternative durable options such as the ACE2-based experimental therapies superior to mAbs to address this unprecedented evolution of Omicron virus. In addition, this type of unique ACE2-based virus-trapping molecules can counter all zoonotic SARS coronaviruses, either from unknown animal hosts or from established wild-life reservoirs of SARS-CoV-2, and even seasonal alpha coronavirus NL63 that depends on human ACE2 for infection.
Full article
(This article belongs to the Special Issue Molecular Epidemiology of SARS-CoV-2, 3rd Edition)
►▼
Show Figures
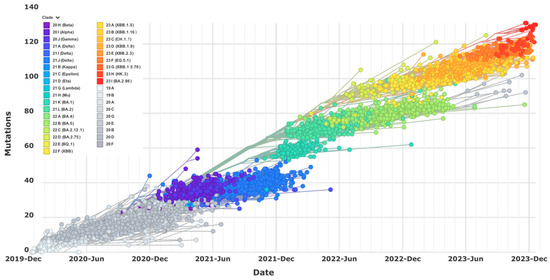
Figure 1
Open AccessArticle
Resistance Mutation Patterns among HIV-1-Infected Children and Features of the Program for Prevention of Mother-to-Child Transmission in Vietnam’s Central Highlands and Southern Regions, 2017–2021
by
Huynh Hoang Khanh Thu, Alexandr N. Schemelev, Yulia V. Ostankova, Diana E. Reingardt, Vladimir S. Davydenko, Nguyen Tuong Vi, Le Ngoc Tu, Ton Tran, Truong Thi Xuan Lien, Aleksandr V. Semenov and Areg A. Totolian
Viruses 2024, 16(5), 696; https://doi.org/10.3390/v16050696 (registering DOI) - 28 Apr 2024
Abstract
The Vietnam Ministry of Health (MOH) has intensified efforts in its aim to eliminate AIDS by 2030. Expanding the program for prevention of mother-to-child transmission (PMTCT) is a significant step towards achieving this goal. However, there are still HIV-exposed children who do not
[...] Read more.
The Vietnam Ministry of Health (MOH) has intensified efforts in its aim to eliminate AIDS by 2030. Expanding the program for prevention of mother-to-child transmission (PMTCT) is a significant step towards achieving this goal. However, there are still HIV-exposed children who do not have access to PMTCT services, and some who have participated in the program but still contracted HIV. This study focused on assessing the prevalence and profile of HIV mutations among children under 18 months of age who had recently tested positive for HIV, while gaining insights into the implementation of early infant diagnostic (EID) tests. Between 2017 and 2021, 3.43% of 5854 collected dry blood spot (DBS) specimens from Vietnam’s Central and Southern regions showed positive EID results. This study identified a high prevalence of resistance mutations in children, totaling 62.9% (95% CI: 53.5–72.3). The highest prevalence of mutations was observed for NNRTIs, with 57.1% (95% CI: 47.5–66.8). Common mutations included Y181C and K103N (NNRTI resistance), M184I/V (NRTI resistance), and no major mutations for PI. The percentage of children with any resistance mutation was significantly higher among those who received PMTCT interventions (69.2%; 95% CI: 50.5–92.6%) compared with those without PMTCT (45.0%; 95% CI: 26.7–71.1%) with χ2 = 6.06, p = 0.0138, and OR = 2.75 (95% CI: 1.13–6.74). Mutation profiles revealed that polymorphic mutations could be present regardless of whether PMTCT interventions were implemented or not. However, non-polymorphic drug resistance mutations were predominantly observed in children who received PMTCT measures. Regarding PMTCT program characteristics, this study highlights the issue of late access to HIV testing for both mothers and their infected children. Statistical differences were observed between PMTCT and non-PMTCT children. The proportion of late detection of HIV infection and breastfeeding rates were significantly higher among non-PMTCT children (p < 0.05). Comparative analysis between children with low viral load (≤200 copies/mL) and high viral load (>200 copies/mL) showed significant differences between the mothers’ current ART regimens (p = 0.029) and the ARV prophylaxis regimen for children (p = 0.016). These findings emphasize the need for comprehensive surveillance to assess the effectiveness of the PMTCT program, including potential transmission of HIV drug-resistance mutations from mothers to children in Vietnam.
Full article
(This article belongs to the Special Issue HIV Reservoirs, Latency, and the Factors Responsible)
►▼
Show Figures
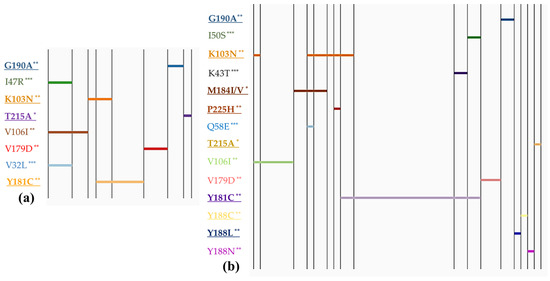
Figure 1
Open AccessArticle
Factor-Dependent Internal Ribosome Entry Site and -1 Programmed Frameshifting Signal in the Bemisia-Associated Dicistrovirus 2
by
Yihang Chen, Subash Chapagain, Jodi Chien, Higor Sette Pereira, Trushar R. Patel, Alice K. Inoue-Nagata and Eric Jan
Viruses 2024, 16(5), 695; https://doi.org/10.3390/v16050695 (registering DOI) - 28 Apr 2024
Abstract
The dicistrovirus intergenic (IGR) IRES uses the most streamlined translation initiation mechanism: the IRES recruits ribosomes directly without using protein factors and initiates translation from a non-AUG codon. Several subtypes of dicistroviruses IRES have been identified; typically, the IRESs adopt two -to three
[...] Read more.
The dicistrovirus intergenic (IGR) IRES uses the most streamlined translation initiation mechanism: the IRES recruits ribosomes directly without using protein factors and initiates translation from a non-AUG codon. Several subtypes of dicistroviruses IRES have been identified; typically, the IRESs adopt two -to three overlapping pseudoknots with key stem-loop and unpaired regions that interact with specific domains of the ribosomal 40S and 60S subunits to direct translation. We previously predicted an atypical IGR IRES structure and a potential -1 programmed frameshift (-1 FS) signal within the genome of the whitefly Bemisia-associated dicistrovirus 2 (BaDV-2). Here, using bicistronic reporters, we demonstrate that the predicted BaDV-2 -1 FS signal can drive -1 frameshifting in vitro via a slippery sequence and a downstream stem-loop structure that would direct the translation of the viral RNA-dependent RNA polymerase. Moreover, the predicted BaDV-2 IGR can support IRES translation in vitro but does so through a mechanism that is not typical of known factorless dicistrovirus IGR IRES mechanisms. Using deletion and mutational analyses, the BaDV-2 IGR IRES is mapped within a 140-nucleotide element and initiates translation from an AUG codon. Moreover, the IRES does not bind directly to purified ribosomes and is sensitive to eIF2 and eIF4A inhibitors NSC1198983 and hippuristanol, respectively, indicating an IRES-mediated factor-dependent mechanism. Biophysical characterization suggests the BaDV-2 IGR IRES contains several stem-loops; however, mutational analysis suggests a model whereby the IRES is unstructured or adopts distinct conformations for translation initiation. In summary, we have provided evidence of the first -1 FS frameshifting signal and a novel factor-dependent IRES mechanism in this dicistrovirus family, thus highlighting the diversity of viral RNA-structure strategies to direct viral protein synthesis.
Full article
(This article belongs to the Special Issue Viral Strategies and Cellular Countermeasures that Regulate mRNA Access to the Translation Apparatus)
►▼
Show Figures
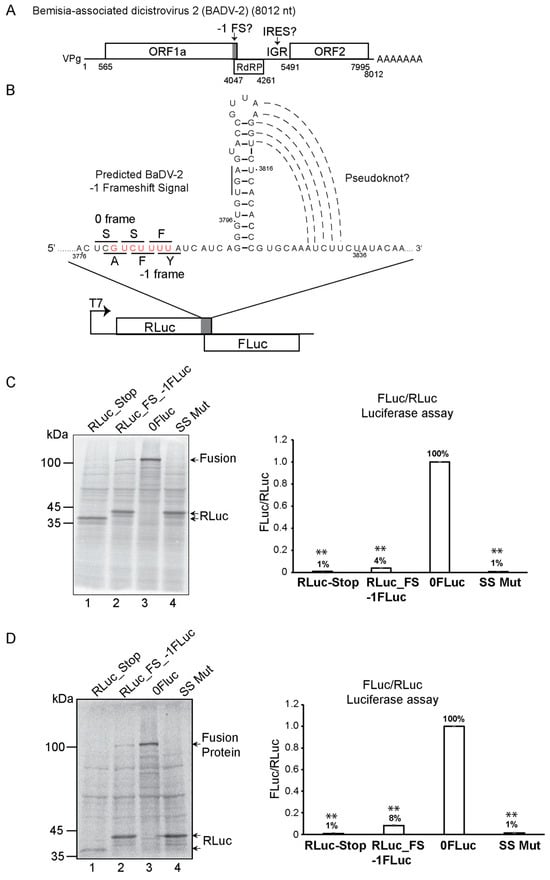
Figure 1
Open AccessArticle
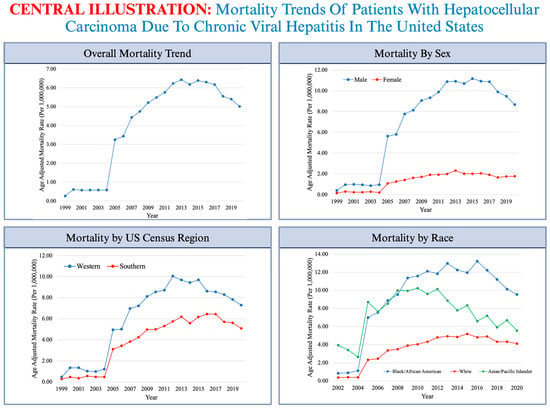
A Longitudinal Analysis of Mortality Related to Chronic Viral Hepatitis and Hepatocellular Carcinoma in the United States
by
N. Begum Ozturk, Hoang Nhat Pham, Rama Mouhaffel, Ramzi Ibrahim, Marwan Alsaqa, Ahmet Gurakar and Behnam Saberi
Viruses 2024, 16(5), 694; https://doi.org/10.3390/v16050694 (registering DOI) - 28 Apr 2024
Abstract
(1) Background: Hepatocellular carcinoma (HCC) contributes to the significant burden of cancer mortality in the United States (US). Despite highly efficacious antivirals, chronic viral hepatitis (CVH) remains an important cause of HCC. With advancements in therapeutic modalities, along with the aging of the
[...] Read more.
(1) Background: Hepatocellular carcinoma (HCC) contributes to the significant burden of cancer mortality in the United States (US). Despite highly efficacious antivirals, chronic viral hepatitis (CVH) remains an important cause of HCC. With advancements in therapeutic modalities, along with the aging of the population, we aimed to assess the contribution of CVH in HCC-related mortality in the US between 1999–2020. (2) Methods: We queried all deaths related to CVH and HCC in the multiple-causes-of-death files from the CDC Wide-ranging Online Data for Epidemiologic Research (WONDER) database between 1999–2020. Using the direct method of standardization, we adjusted all mortality information for age and compared the age-adjusted mortality rates (AAMRs) across demographic populations and by percentile rankings of social vulnerability. Temporal shifts in mortality were quantified using log-linear regression models. (3) Results: A total of 35,030 deaths were identified between 1999–2020. The overall crude mortality increased from 0.27 in 1999 to 8.32 in 2016, followed by a slight reduction to 7.04 in 2020. The cumulative AAMR during the study period was 4.43 (95% CI, 4.39–4.48). Males (AAMR 7.70) had higher mortality rates compared to females (AAMR 1.44). Mortality was higher among Hispanic populations (AAMR 6.72) compared to non-Hispanic populations (AAMR 4.18). Higher mortality was observed in US counties categorized as the most socially vulnerable (AAMR 5.20) compared to counties that are the least socially vulnerable (AAMR 2.53), with social vulnerability accounting for 2.67 excess deaths per 1,000,000 person-years. (4) Conclusions: Our epidemiological analysis revealed an overall increase in CVH-related HCC mortality between 1999–2008, followed by a stagnation period until 2020. CVH-related HCC mortality disproportionately affected males, Hispanic populations, and Black/African American populations, Western US regions, and socially vulnerable counties. These insights can help aid in the development of strategies to target vulnerable patients, focus on preventive efforts, and allocate resources to decrease HCC-related mortality.
Full article
(This article belongs to the Section Human Virology and Viral Diseases)
►▼
Show Figures
Graphical abstract
Open AccessArticle
EcoHIV Infection of Primary Murine Brain Cell Cultures to Model HIV Replication and Neuropathogenesis
by
Boe-Hyun Kim, Wei Chao, Eran Hadas, Alejandra Borjabad, Mary Jane Potash and David J. Volsky
Viruses 2024, 16(5), 693; https://doi.org/10.3390/v16050693 (registering DOI) - 27 Apr 2024
Abstract
Background. EcoHIV is a chimeric HIV that replicates in mice in CD4+ T cells, macrophages, and microglia (but not in neurons), causing lasting neurocognitive impairment resembling neurocognitive disease in people living with HIV. The present study was designed to develop EcoHIV-susceptible primary mouse
[...] Read more.
Background. EcoHIV is a chimeric HIV that replicates in mice in CD4+ T cells, macrophages, and microglia (but not in neurons), causing lasting neurocognitive impairment resembling neurocognitive disease in people living with HIV. The present study was designed to develop EcoHIV-susceptible primary mouse brain cultures to investigate the indirect effects of HIV infection on neuronal integrity. Results. We used two EcoHIV clones encoding EGFP and mouse bone marrow-derived macrophages (BMM), mixed mouse brain cells, or enriched mouse glial cells from two wild-type mouse strains to test EcoHIV replication efficiency, the identity of productively infected cells, and neuronal apoptosis and integrity. EcoHIV replicated efficiently in BMM. In mixed brain cell cultures, EcoHIV targeted microglia but did not cause neuronal apoptosis. Instead, the productive infection of the microglia activated them and impaired synaptophysin expression, dendritic density, and axonal structure in the neurons. EcoHIV replication in the microglia and neuronal structural changes during infection were prevented by culture with an antiretroviral. Conclusions. In murine brain cell cultures, EcoHIV replication in the microglia is largely responsible for the aspects of neuronal dysfunction relevant to cognitive disease in infected mice and people living with HIV. These cultures provide a tool for further study of HIV neuropathogenesis and its control.
Full article
(This article belongs to the Special Issue Roles of Macrophages in Viral Infections)
►▼
Show Figures
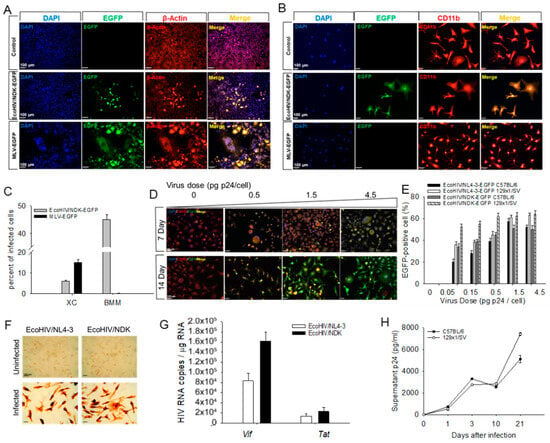
Figure 1
Open AccessBrief Report
Influenza Virus Genomic Surveillance, Arizona, USA, 2023–2024
by
Rabia Maqsood, Matthew F. Smith, LaRinda A. Holland, Regan A. Sullins, Steven C. Holland, Michelle Tan, Gabrielle M. Hernandez Barrera, Alexis W. Thomas, Mario Islas, Joanna L. Kramer, Lora Nordstrom, Mary Mulrow, Michael White, Vel Murugan and Efrem S. Lim
Viruses 2024, 16(5), 692; https://doi.org/10.3390/v16050692 (registering DOI) - 27 Apr 2024
Abstract
Influenza viruses are constantly evolving and are therefore monitored worldwide in the hope to reduce the burden of disease by annual updates to vaccine recommendations. We conducted genomic sequencing of 110 influenza A and 30 influenza B viruses from specimens collected between October
[...] Read more.
Influenza viruses are constantly evolving and are therefore monitored worldwide in the hope to reduce the burden of disease by annual updates to vaccine recommendations. We conducted genomic sequencing of 110 influenza A and 30 influenza B viruses from specimens collected between October 2023 and February 2024 in Arizona, USA. We identified mutations in the hemagglutinin (HA) antigenic sites as well as the neuraminidase (NA) gene in our samples. We also found no unique HA and NA mutations in vaccinated yet influenza-infected individuals. Real-time genomic sequencing surveillance is important to ensure influenza vaccine effectiveness.
Full article
(This article belongs to the Special Issue Influenza and Other Respiratory Viruses: Prevention, Diagnosis, Treatment)
►▼
Show Figures
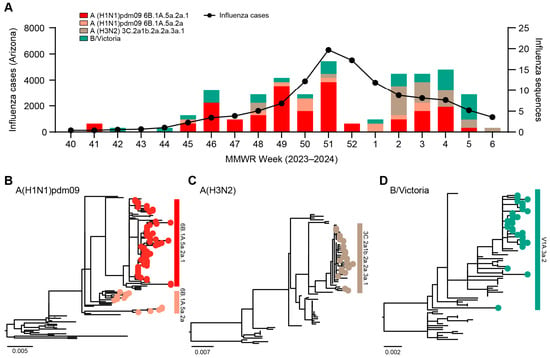
Figure 1
Open AccessArticle
Diagnostic Utility of Cytomegalovirus (CMV) DNA Quantitation in Ulcerative Colitis
by
Sema Esen, Imran Saglik, Enver Dolar, Selcan Cesur, Nesrin Ugras, Harun Agca, Osman Merdan and Beyza Ener
Viruses 2024, 16(5), 691; https://doi.org/10.3390/v16050691 (registering DOI) - 26 Apr 2024
Abstract
Cytomegalovirus (CMV) colitis is a critical condition associated with severe complications in ulcerative colitis (UC). This study aimed to investigate the diagnostic value of the presence of CMV DNA in intestinal mucosa tissue and blood samples in patients with active UC. This study
[...] Read more.
Cytomegalovirus (CMV) colitis is a critical condition associated with severe complications in ulcerative colitis (UC). This study aimed to investigate the diagnostic value of the presence of CMV DNA in intestinal mucosa tissue and blood samples in patients with active UC. This study included 81 patients with exacerbated symptoms of UC. Patient data were obtained from the Hospital Information Management System. CMV DNA in colorectal tissue and plasma samples were analyzed using a real-time quantitative PCR assay. CMV markers were detected using immunohistochemistry and hematoxylin–eosin staining. Immunohistochemistry positivity was observed in tissue samples from eight (9.9%) patients. Only one (1.2%) patient showed CMV-specific intranuclear inclusion bodies. CMV DNA was detected in 63.0% of the tissues (median: 113 copies/mg) and in 58.5% of the plasma samples (median: 102 copies/mL). For tissues, sensitivity and the negative predictive value (NPV) for qPCR were excellent (100.0%), whereas specificity and the positive predictive value (PPV) were low (41.9% and 15.7%, respectively). For plasma, sensitivity and NPV were high (100.0%) for qPCR, whereas specificity and PPV were low (48.6% and 24.0%, respectively). CMV DNA ≥392 copies/mg in tissue samples (sensitivity 100.0% and specificity 83.6%) and ≥578 copies/mL (895 IU/mL) in plasma samples (sensitivity 66.7% and specificity 100.0%) provided an optimal diagnosis for this test. The qPCR method improved patient management through the early detection of CMV colitis in patients with UC. However, reliance on qPCR positivity alone can lead to overdiagnosis. Quantification of CMV DNA can improve diagnostic specificity, although standardization is warranted.
Full article
(This article belongs to the Section Human Virology and Viral Diseases)

Journal Menu
► ▼ Journal Menu-
- Viruses Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Diseases, Infectious Disease Reports, Pathogens, Viruses, TropicalMed
Human Monkeypox Research
Topic Editors: Shailendra K. Saxena, Ahmed Sayed Abdel-MoneimDeadline: 30 June 2024
Topic in
Biomedicines, JCM, Pathogens, Vaccines, Viruses
Discovery and Development of Monkeypox Disease Treatments
Topic Editors: Mohd Imran, Ali A. RabaanDeadline: 31 August 2024
Topic in
Brain Sciences, Clinics and Practice, COVID, Life, Vaccines, Viruses
Multifaceted Efforts from Basic Research to Clinical Practice in Controlling COVID-19 Disease
Topic Editors: Yih-Horng Shiao, Rashi OjhaDeadline: 30 September 2024

Conferences
Special Issues
Special Issue in
Viruses
Oncolytic Viruses as Immunotherapeutic Agents
Guest Editor: Nadine Van MontfoortDeadline: 1 May 2024
Special Issue in
Viruses
Animal Coronaviruses: Infection, Prevention, and Antivirals
Guest Editor: Tomomi TakanoDeadline: 20 May 2024
Special Issue in
Viruses
Bacteriophages and Biofilms 2.0
Guest Editors: Zuzanna Drulis-Kawa, Tomasz OlszakDeadline: 31 May 2024
Special Issue in
Viruses
The Inflammasomes - Key Players in Antiviral Response
Guest Editors: Dong-Yan Jin, Tsan Sam XiaoDeadline: 15 June 2024
Topical Collections
Topical Collection in
Viruses
Poxviruses
Collection Editors: Giliane de Souza Trindade, Galileu Barbosa Costa, Flavio Guimaraes da Fonseca
Topical Collection in
Viruses
Phage Therapy
Collection Editors: Nina Chanishvili, Jean-Paul Pirnay, Mikael Skurnik
Topical Collection in
Viruses
Coronaviruses
Collection Editors: Luis Martinez-Sobrido, Fernando Almazan Toral
Topical Collection in
Viruses
SARS-CoV-2 and COVID-19
Collection Editors: Luis Martinez-Sobrido, Fernando Almazan Toral